 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK10560.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 299aa MW: 34613.5 Da PI: 9.3178 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 18.8 | 4.5e-06 | 23 | 47 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp dC+ +++rk++L+rH+ H
PK10560.1 23 HVCPvdDCPSRYRRKDHLNRHLLIH 47
79********************998 PP
| |||||||
| 2 | zf-C2H2 | 16.3 | 2.8e-05 | 95 | 119 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F +s L++H ++H
PK10560.1 95 HVCQeaGCGKAFAFPSRLRKHEQSH 119
789999***************9998 PP
| |||||||
| 3 | zf-C2H2 | 14.5 | 0.0001 | 187 | 210 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C C +Fs+ksnL++H++ H
PK10560.1 187 CLykGCLCTFSTKSNLNQHLKAvH 210
66679***************9877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 9.8E-10 | 20 | 49 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.31E-6 | 20 | 49 | No hit | No description |
| PROSITE profile | PS50157 | 8.995 | 23 | 52 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.004 | 23 | 47 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 25 | 47 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.27E-5 | 50 | 78 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-4 | 50 | 78 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.136 | 52 | 82 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.11 | 52 | 77 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 54 | 77 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.6E-8 | 89 | 119 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.78E-5 | 91 | 121 | No hit | No description |
| PROSITE profile | PS50157 | 10.866 | 95 | 124 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.013 | 95 | 119 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 97 | 119 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 10 | 127 | 152 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 129 | 152 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 155 | 176 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-4 | 166 | 207 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.64E-7 | 166 | 222 | No hit | No description |
| PROSITE profile | PS50157 | 11.365 | 185 | 215 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.029 | 185 | 210 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 187 | 210 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.9E-5 | 209 | 238 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 7.8 | 216 | 242 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 218 | 242 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
XGDLLRSSFK RTLISYIQFE RPHVCPVDDC PSRYRRKDHL NRHLLIHKGK LFKCPIENCK 60 SEFSVKANIK RHVEELHNEG RPSPSTECEN TERKHVCQEA GCGKAFAFPS RLRKHEQSHV 120 KLESIEALCA EPGCMKVFTN EQCLKGHIEL CHRHISCKVC GSKHLKKNMK RHLRTHERKS 180 SFESIKCLYK GCLCTFSTKS NLNQHLKAVH FNQKPYVCGF SGCGMRFAYK HVRDNHEKSS 240 CHVYTHGDFE EADEQFRSRP RGGRKRKYPS LVELLVRKRI TPPNDLDGLF ECSDNLSCG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 164 | 170 | LKKNMKR |
| 2 | 258 | 266 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
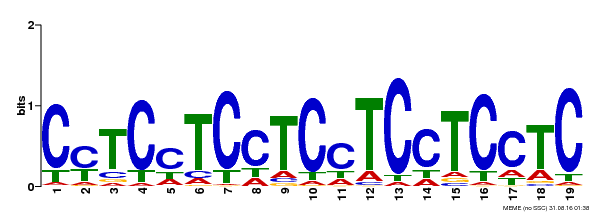
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010104482.1 | 1e-136 | transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 1e-113 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A2P5E9Z9 | 1e-148 | A0A2P5E9Z9_TREOI; TFIIH C1-like domain containing protein | ||||
| STRING | XP_010104482.1 | 1e-136 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3069 | 33 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.1 | 1e-116 | transcription factor IIIA | ||||



