 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK08504.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 841aa MW: 91400.2 Da PI: 5.8755 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.7e-21 | 124 | 179 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
PK08504.1 124 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 179
688999***********************************************999 PP
| |||||||
| 2 | START | 207.2 | 6.4e-65 | 330 | 555 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECT CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetleviss 87
ela++a++elvk+a+ eep+W+ks e +n++e++++f + + + +ea+r+sg+v+ ++ lve+l++++ +W e+++ + +t++viss
PK08504.1 330 ELALAAMDELVKMAQTEEPLWMKSFeggrEVLNEEEYMRSFNPCIGmkpngFVTEASRESGIVIINSLALVETLMESS-RWAEMFPcviaRTSTTDVISS 428
5899*********************9**9************9988899999***************************.********************* PP
T......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHH CS
START 88 g......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphw 179
g galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d +++++ +s++ +++lpSg+++++++ g+skvtwveh++++++++h+
PK08504.1 429 GmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRDNSSgPPSFINCRRLPSGCVVQDMPSGYSKVTWVEHAEYDENQIHQ 528
*********************************************************999**************************************** PP
HHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 180 llrslvksglaegaktwvatlqrqcek 206
l+r+l++sg+ +ga++w atlqrqce+
PK08504.1 529 LYRPLLSSGMGFGAQRWIATLQRQCEC 555
*************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.03E-20 | 106 | 181 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.5E-22 | 111 | 181 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 121 | 181 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 122 | 185 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.38E-18 | 123 | 181 | No hit | No description |
| Pfam | PF00046 | 1.4E-18 | 124 | 179 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 156 | 179 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.961 | 321 | 558 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.94E-33 | 323 | 555 | No hit | No description |
| CDD | cd08875 | 1.21E-122 | 325 | 554 | No hit | No description |
| SMART | SM00234 | 5.9E-49 | 330 | 555 | IPR002913 | START domain |
| Pfam | PF01852 | 1.7E-57 | 330 | 555 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.94E-23 | 584 | 754 | No hit | No description |
| SuperFamily | SSF55961 | 3.94E-23 | 794 | 834 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 841 aa Download sequence Send to blast |
MSFGGFLDNS TSTSGGGARL VAEIPYSSTS NNDNNHIMPS TAIAQPRLIT QSLTKSMFNS 60 PGLSLALQTN IDGQGDMIRN MAESFEPGSG RRSRDEEHES RSGSDNMDGG SGDDQDATDK 120 PPRKKRYHRH TPQQIQELEA LFKECPHPDE KQRLELSKRL CLETRQVKFW FQNRRTQMKT 180 QLERHENSLL RQENDKLRAE NMSIRDAMRN PMCTNCGGPA IIGEISLEEQ HLRIENARLK 240 DELERVCSLA GKFLGRPISA LATSMAPPLP SSALELGVGS NGFGGLTSVS TTMPDFGISS 300 NTLAALPPQS RSQGGVQVLD RSIERSMYLE LALAAMDELV KMAQTEEPLW MKSFEGGREV 360 LNEEEYMRSF NPCIGMKPNG FVTEASRESG IVIINSLALV ETLMESSRWA EMFPCVIART 420 STTDVISSGM GGTRNGALQL MHAELQVLSP LVPVREVNFL RFCKQHAEGV WAVVDVSIDT 480 IRDNSSGPPS FINCRRLPSG CVVQDMPSGY SKVTWVEHAE YDENQIHQLY RPLLSSGMGF 540 GAQRWIATLQ RQCECLAILM SSTVPTRDHT AGITSSGRRS MLKLAQRMTD NFCAGVCAST 600 VHKWNKLNAG NVDEDVRVMT RKSVDDPGEP PGIVLSAATS VWLPVSPQRL FDFLRDESLR 660 SEWDILSNGG PMQEMAHIAK GQNHGNCVSL LRASAMNTNQ SSMLILQETC IDEAGSLVVY 720 APVDIPAMHV VMNGGDSAYV ALLPSGFAIV PDGPGSRGPN LGSTPNGGCS NNNNNNGGSG 780 SNGNGGGDGS HRASGSLLTV AFQILVNSLP TAKLTVESVE TVNNLISCTV QKIKAALQCE 840 S |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
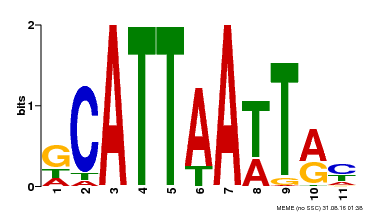
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024018220.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2P5CNQ4 | 0.0 | A0A2P5CNQ4_TREOI; Octamer-binding transcription factor | ||||
| STRING | XP_010091553.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



