 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK02936.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 272aa MW: 30930.9 Da PI: 8.2053 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 103.8 | 1e-32 | 18 | 72 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LH+rFv+av++LGG+ekAtPk++l+lm++kgLtl+h+kSHLQkYRl
PK02936.2 18 KPRLRWTADLHDRFVDAVTKLGGPEKATPKAVLRLMGLKGLTLYHLKSHLQKYRL 72
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.538 | 15 | 75 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-31 | 15 | 73 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.9E-24 | 18 | 73 | IPR006447 | Myb domain, plants |
| SuperFamily | SSF46689 | 7.88E-16 | 18 | 75 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.3E-9 | 20 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.1E-22 | 118 | 164 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
MERSFGYENG IVMTRDPKPR LRWTADLHDR FVDAVTKLGG PEKATPKAVL RLMGLKGLTL 60 YHLKSHLQKY RLGQQTRKQN LVEHSKESNG CSYVQFGSHS SVTSTTSSRG DKEQCDIPIT 120 EALKHQNEVQ KRLEEQLEVQ KKLQMRIEAQ GKYLQAILEK AQKSLSLDMN GPGNIEVAKA 180 QLTDFNLALS NFMENMTEVE EKASIIDQMG GVYRKQNDSS FKIYQESNRE DSEDSKLKDT 240 NTIHFDLNTK GGYDFVGANS LEFEPKMLSY GR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 6e-19 | 18 | 71 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-19 | 18 | 71 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-19 | 18 | 71 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-19 | 18 | 71 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 7e-19 | 18 | 71 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
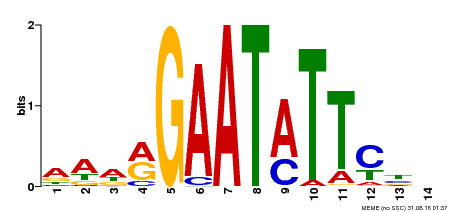
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010110423.2 | 1e-150 | myb family transcription factor PHL11 | ||||
| Swissprot | C0SVS4 | 1e-86 | PHLB_ARATH; Myb family transcription factor PHL11 | ||||
| TrEMBL | A0A2P5FA92 | 1e-164 | A0A2P5FA92_TREOI; Octamer-binding transcription factor | ||||
| STRING | XP_010110423.1 | 1e-149 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5101 | 31 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 6e-84 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



