 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10023478m | ||||||||
| Common Name | CARUB_v10023478mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 368aa MW: 39835.8 Da PI: 7.9786 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 137.3 | 4.4e-43 | 72 | 149 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC +dl++ak y++rh+vC vhsk+p+v+v+gleqrfCqqCsrfh+l efD ekrsCrrrLa+hnerrrk+q+
Carubv10023478m 72 RCQVEGCGMDLTNAKGYYSRHRVCGVHSKTPKVIVAGLEQRFCQQCSRFHQLPEFDLEKRSCRRRLAGHNERRRKPQP 149
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-34 | 65 | 134 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.017 | 70 | 147 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.19E-40 | 71 | 151 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-32 | 73 | 146 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:2000025 | Biological Process | regulation of leaf formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 368 aa Download sequence Send to blast |
MEMGSNSGPG LGPGQAESGG SSTESSSFSG GLMFGQKIYF EDGGGGSGSS SSGGSNRRVR 60 GGGSGQSGQI PRCQVEGCGM DLTNAKGYYS RHRVCGVHSK TPKVIVAGLE QRFCQQCSRF 120 HQLPEFDLEK RSCRRRLAGH NERRRKPQPA SLSVLSSRYG RIAPSLYGNA DAGMNGSFLG 180 NQDMGWTSAR TLDTRVMRRP VSAPSWQISP MNVFSQGSVG GGGTSFSSPE IMDTKLESYK 240 GMGDSNCALS LLSNPHQPQD NNNNNNTWRA SSGFCPMTVT MAQPPPAPGQ HQYLNPPWVF 300 KDDDSDMSPV LNLGRFTEPD NCQISNGTTM GEFELSDHHH QSRRQYIEDE QNTRAYDSSS 360 HHNNWSL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 7e-31 | 62 | 146 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
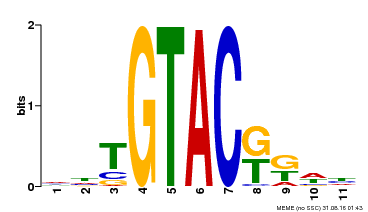
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00307 | DAP | Transfer from AT2G42200 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10023478m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156 and miR157. {ECO:0000305|PubMed:12202040}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353538 | 0.0 | AK353538.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-50-J24. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006294465.1 | 0.0 | squamosa promoter-binding-like protein 9 | ||||
| Swissprot | Q700W2 | 0.0 | SPL9_ARATH; Squamosa promoter-binding-like protein 9 | ||||
| TrEMBL | R0HTJ6 | 0.0 | R0HTJ6_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006294465.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2144 | 27 | 75 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42200.1 | 0.0 | squamosa promoter binding protein-like 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10023478m |
| Entrez Gene | 17887996 |



