 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10009866m | ||||||||
| Common Name | CARUB_v10009866mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 303aa MW: 32876.6 Da PI: 7.1892 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 50.9 | 2.2e-16 | 223 | 258 | 1 | 34 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkg 34
C++Cg ++ Tp++Rrgp+g++tLCnaCGl+++ kg
Carubv10009866m 223 CRHCGISEksTPMMRRGPEGPRTLCNACGLMWANKG 258
*****99999***********************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 14.513 | 77 | 112 | IPR010399 | Tify domain |
| SMART | SM00979 | 7.6E-11 | 77 | 112 | IPR010399 | Tify domain |
| Pfam | PF06200 | 3.0E-11 | 79 | 111 | IPR010399 | Tify domain |
| Pfam | PF06203 | 2.1E-15 | 147 | 189 | IPR010402 | CCT domain |
| PROSITE profile | PS51017 | 12.987 | 147 | 189 | IPR010402 | CCT domain |
| SMART | SM00401 | 2.6E-12 | 217 | 270 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 9.156 | 217 | 265 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 4.66E-13 | 218 | 282 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.3E-15 | 221 | 264 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 5.47E-15 | 222 | 272 | No hit | No description |
| Pfam | PF00320 | 2.8E-14 | 223 | 258 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 223 | 250 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 303 aa Download sequence Send to blast |
MDDLHGSNAR MHIGEAQDSM HVQYEHHALH HIHNGSGMVD DHADGGNAGG MSEGVETDIP 60 SHPGNITDNR GEVVDRGSEQ GDQLTLSFQG QVYVFDSVLP EKVQAVLLLL GGRELPQATP 120 TGLGSSHHNN RALGLPGTPQ RFSIPQRLAS LVRFREKRKG RNFDKKIRYT VRKEVALRMQ 180 RNKGQFTSAK SVNDEAASAG SSWGSNQTWA IEGSEAQNQE ISCRHCGISE KSTPMMRRGP 240 EGPRTLCNAC GLMWANKGAL RDLSKGAPPQ TPQNVPLSKN EDANLEADQM MISVANNISN 300 SQ* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
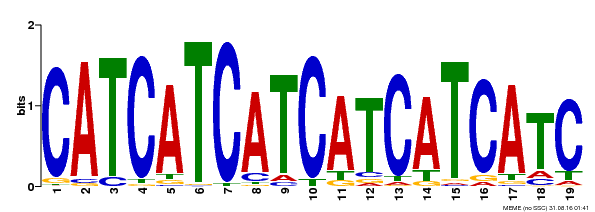
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10009866m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB119061 | 0.0 | AB119061.1 Arabidopsis thaliana ZML2 mRNA for GATA-type zinc finger protein, complete cds. | |||
| GenBank | AY150395 | 0.0 | AY150395.1 Arabidopsis thaliana putative flowering protein CONSTANS (At1g51600) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006305455.1 | 0.0 | GATA transcription factor 28 isoform X1 | ||||
| Refseq | XP_023644658.1 | 0.0 | GATA transcription factor 28 isoform X1 | ||||
| Swissprot | Q8H1G0 | 0.0 | GAT28_ARATH; GATA transcription factor 28 | ||||
| TrEMBL | R0IMP1 | 0.0 | R0IMP1_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006305455.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3684 | 26 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51600.2 | 0.0 | ZIM-LIKE 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10009866m |
| Entrez Gene | 17899194 |



