 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10004076m | ||||||||
| Common Name | CARUB_v10004076mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 960aa MW: 105622 Da PI: 5.7443 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.9 | 4.9e-29 | 590 | 640 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
Carubv10004076m 590 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 640
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF55781 | 3.63E-5 | 213 | 284 | IPR029016 | GAF domain-like |
| PROSITE profile | PS51519 | 17.757 | 579 | 660 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.3E-26 | 592 | 640 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.27E-23 | 860 | 948 | No hit | No description |
| SMART | SM00666 | 2.4E-28 | 863 | 945 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 27.793 | 863 | 945 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.0E-27 | 863 | 945 | No hit | No description |
| Pfam | PF00564 | 3.1E-20 | 863 | 944 | IPR000270 | PB1 domain |
| CDD | cd06407 | 4.18E-34 | 864 | 944 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 960 aa Download sequence Send to blast |
MCEPDDNSAR NGVAPPPSRS RELLMDVDDL DLDGSWPLDQ IPYLSSSNRM ISPIFVSSSS 60 EQPCSPLWAF SDGGGNGNHH HANAGADDDK ISSASVVPSF RLADYPLFLP YSSPSAAENT 120 TEKHNNFQFP SPLMSLVPPE NTDNYCVIKE RMTQALRYFK DSTEQHVLAQ VWAPVRKNGR 180 DLLTTLGQPF VLNPNGNGLN QYRMISLTYM FSVDSESDIE LGLPGRVFRQ KLPEWTPNVQ 240 YYSSKEFSRL DHALHYNVRG TLALPVFNPS GQSCIGVVEL IMTSEKIHYA PEVDKVCKAL 300 EAVNLKSSEI LDHQTTQICN ESRQNALAEI LEVLTVVCET HNLPLAQTWV PCQHGSVLAN 360 GGGLKKNCTS FDGSCMGQIC MSTTDMACYV VDAHVWGFRD ACLEHHLQKG QGVAGRAFLN 420 GGSCFCRDIT KFCKTQYPLV HYALMFKLTT CFAISLQSSY TGDDSYILEF FLPSSITDDQ 480 EQDSLLGSIL VTMKEHFQSL RVASGVDFGE DDDKLSFEII QALPDTKIHS KIESIRVPFS 540 GFKSNATETR VIPQPVVQSS DPVNENGNVA TVNGVVKEKK KTEKKRGKTE KTISLDVLQQ 600 YFTGSLKDAA KSLGVCPTTM KRICRQHGIS RWPSRKIKKV NRSITKLKRV IESVQGTDGG 660 LDLTSMAVSS IPWTHGQPSA QPLNSPSGSK PPELPNANNS PNHWSSDHSP HEPNCSPELP 720 SNGHKRSRTV DESAGTPTSH GSCDGNQLDE TKVPNQDPLF TVGESPGLLF PPYARDHDVS 780 AASFSMPNRL LGSIDHFRGM LIEDAGSSKD LRNLCSTAAF DDKFPDSNWM NNDNNSNNNI 840 YAPPKEEAIA NVTRGASGSE TRTITIKASY KEDIIRFRIS SGSGIMELKD EVAKRLKLDA 900 GTFDIKYLDD DNEWVLIACD ADLQECLEIP RSSRTNIVRL LVHDVTTNLG SSCESTGEL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
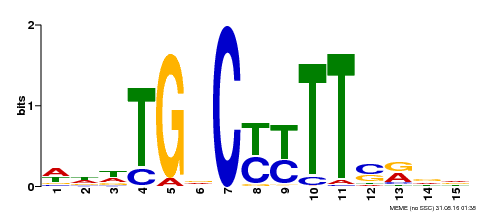
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10004076m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228452 | 0.0 | AK228452.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-03-P13. | |||
| GenBank | BT005776 | 0.0 | BT005776.1 Arabidopsis thaliana clone RAFL15-03-P13 (R20174) unknown protein (At4g24020) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006282537.1 | 0.0 | protein NLP7 | ||||
| Refseq | XP_023634148.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | R0GSX5 | 0.0 | R0GSX5_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006282537.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10004076m |
| Entrez Gene | 17881043 |



