 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10003148m | ||||||||
| Common Name | CARUB_v10003148mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 342aa MW: 38321.9 Da PI: 6.0951 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 59.9 | 5.4e-19 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ede+lv++v+++G +W++ ++ g++R++k+c++rw +yl
Carubv10003148m 14 KGPWTPDEDEKLVNYVQEHGHSSWRALPKLAGLNRCGKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.8 | 4.6e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgr++++E++ +++++ lG++ W+tIa+ ++ gRt++++k++w+++l
Carubv10003148m 67 RGRFSPDEEQTILNLHSVLGNK-WSTIANQLP-GRTDNEIKNFWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-24 | 6 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.088 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.47E-31 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.0E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.5E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.91E-12 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.031 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-27 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.3E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.5E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.73E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MGRPPSSDDS GLKKGPWTPD EDEKLVNYVQ EHGHSSWRAL PKLAGLNRCG KSCRLRWTNY 60 LRPDIKRGRF SPDEEQTILN LHSVLGNKWS TIANQLPGRT DNEIKNFWNT HLKKKLIQMG 120 IDPMTHRPRT DIFSGLSQLM SMSSNLRGFA ELQQQFPISQ EHTILRLQTE MAKLQLFQYL 180 LQPSSFMNTI NPNDLDSLSL LNSIASFKET TTTTTSNTLD LGCLGSYLQD FNSLPSLKTL 240 NSNSMEPSSV FPQNPDDNHF KFFTQRQNLP VSPIWLSDPS SSTQSLTLPN SVVTDDLIRN 300 QYGVEDVNSN ITSSSCQESA ASASAAWPDH LLDDSIFSDI P* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 3e-29 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250|UniProtKB:Q9M2D9}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00503 | DAP | Transfer from AT5G10280 | Download |
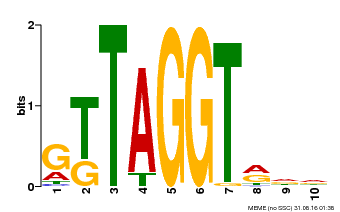
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10003148m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by gravity in roots. {ECO:0000269|PubMed:24902892}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY519619 | 0.0 | AY519619.1 Arabidopsis thaliana MYB transcription factor (At5g10280) mRNA, complete cds. | |||
| GenBank | BT002877 | 0.0 | BT002877.1 Arabidopsis thaliana clone RAFL14-88-D17 (R20400) putative myb family transcription factor (At5g10280) mRNA, complete cds. | |||
| GenBank | BT004402 | 0.0 | BT004402.1 Arabidopsis thaliana clone U20400 putative myb family transcription factor (At5g10280) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006289596.1 | 0.0 | transcription factor MYB41 | ||||
| Refseq | XP_023637704.1 | 0.0 | transcription factor MYB41 | ||||
| Swissprot | Q9SBF3 | 0.0 | MYB92_ARATH; Transcription factor MYB92 | ||||
| TrEMBL | R0GZZ0 | 0.0 | R0GZZ0_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006289596.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10280.1 | 0.0 | myb domain protein 92 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10003148m |
| Entrez Gene | 17882827 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



