 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_86286_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 220aa MW: 24308.5 Da PI: 6.8105 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.8 | 7.5e-20 | 116 | 167 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++y+GVr+++ +gr++AeIrdp+++g r++lg++ t+e+Aa a+++a+ +++g
cra_locus_86286_iso_1_len_792_ver_3 116 TRYRGVRRRP-WGRFAAEIRDPNKRG--SRIWLGTYETPEDAALAYDRAAFQIRG 167
68********.**********96655..************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 4.01E-29 | 116 | 177 | No hit | No description |
| Pfam | PF00847 | 1.6E-14 | 116 | 167 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 24.553 | 117 | 175 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.3E-37 | 117 | 181 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.9E-32 | 117 | 176 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.64E-23 | 117 | 176 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 7.1E-13 | 118 | 129 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.1E-13 | 157 | 177 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 220 aa Download sequence Send to blast |
XSEESSRISN PNDNTFLPTE VNGDHLEFLA TLTEEAGNIS PQLISVELSS EESSRISSPG 60 DNIFISTEAN RGDLGFNYVD NNSPTSTESQ EEDMAGKKKK AGAAARGKNA PTDQWTRYRG 120 VRRRPWGRFA AEIRDPNKRG SRIWLGTYET PEDAALAYDR AAFQIRGAKA LLNFPHLLNC 180 VAQPARVGQR RRAMESSSSS ESETESHGGA RKKMKQETIL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-29 | 118 | 179 | 6 | 67 | ATERF1 |
| 3gcc_A | 2e-29 | 118 | 179 | 6 | 67 | ATERF1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 189 | 214 | RRRAMESSSSSESETESHGGARKKMK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. {ECO:0000269|PubMed:11950980}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00311 | DAP | Transfer from AT2G44840 | Download |
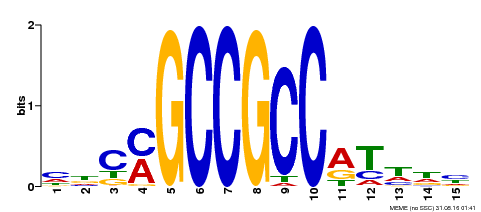
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Strongly induced by wounding. Induced by Pseudomonas syringae tomato (both virulent and avirulent avrRpt2 strains), independently of PAD4. Also induced by methyl jasmonate (MeJA) independently of JAR1, but seems to not be affected by ethylene. Induction by salicylic acid (SA) is controlled by growth and/or developmental conditions, and seems to be more efficient and independent of PAD4 in older plants. {ECO:0000269|PubMed:11950980, ECO:0000269|PubMed:12068110}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027157710.1 | 6e-43 | ethylene-responsive transcription factor 13-like | ||||
| Swissprot | Q8L9K1 | 7e-42 | ERF99_ARATH; Ethylene-responsive transcription factor 13 | ||||
| TrEMBL | A0A192XLN7 | 1e-123 | A0A192XLN7_CATRO; ORCA3-like protein 2 | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA21 | 24 | 1165 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



