 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_778_iso_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 1030aa MW: 113062 Da PI: 5.5282 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.4 | 3.4e-29 | 617 | 668 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
aek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
cra_locus_778_iso_3_len_3340_ver_3 617 AEKTISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 668
589***********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.632 | 607 | 688 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.5E-26 | 620 | 668 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 4.25E-23 | 927 | 1014 | No hit | No description |
| PROSITE profile | PS51745 | 24.855 | 928 | 1010 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 3.2E-19 | 928 | 1009 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.6E-26 | 928 | 1009 | No hit | No description |
| SMART | SM00666 | 1.1E-26 | 928 | 1010 | IPR000270 | PB1 domain |
| CDD | cd06407 | 4.62E-38 | 929 | 1009 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1030 aa Download sequence Send to blast |
MAEPEEPEQQ HQHQKFLHPH PRSMEILTSF QHQPPAERDN IMMDLDLDLD SSWTFDQIFS 60 AAAAAATTTN NNSTSSFLLS TTSEQPCSPL WAFSDENNED KPSGNSTTAL GLRLSDYPRL 120 VNCNNDSITG NPSRNDEKRR FTTPLLELAA LDYPDASCII KERMTQALRF FKDSTESHVL 180 AQVWAPVKNG GRCVLTTSGQ PFVLDPKSNG LYQYRMVSLM YVFSVDGETN GVLGLPGRVF 240 RQKLPEWTPN VQYYSSEEFP RLNHALHYNV RGTLALPVFE PSGQNCIGVL ELIMTSQKIN 300 YAPEVDKVCK ALEAVNLKSS EILDHSNVQI CNEGRQNALA EILEILTVVC ETHKLPLAQT 360 WVPCQHRSVL ANGGGLRKTC SSFDGSCMGQ VCMSTTDVAF YVVDAHMWGF REACAEHHLQ 420 KGQGVAGRAF ESHNSCFCTD VTQFCKNDYP LVHYARMFGL TSCFAICLRS SYTGDDDYVL 480 EFFLPSIVLD HGDQQTLLDS LLVTMKPHLG SLRIASGKDL EHEWRSIEIV KSLVEEKFDS 540 RAEVFQRTRS STSPPGPTAF TNGEMVHLNS MAEQRSALVD SANDERHSGA TAGHQSTASV 600 VENKETGKKT ERKRGKAEKT ISLEVLQQYF AGSLKDAAKS LGVCPTTMKR ICRQHGISRW 660 PSRKINKVNR SLTKLKRVIE SVQGAEGAFT LTSLAASSIP VAVGSVPWTA GGLNESSPQD 720 QPGSGPSEFP GDRNDESPLP GSGSDGKAET SNQLLGGGTG ENEEHVSKSA HFLPCEGSHR 780 SKTQSGSREE SAGTPTSQGS CQGSPYIGNE SSPRNELVVS PSQEQLMKVG GSMEFRYQPT 840 GAINLSAAFS MDAFMNVQHQ EPFGGMLVED AGSCHDLKNL CSNGDALFDE RLPDYSWNNP 900 PSSEPIPKGS TTPADRMPQF SARPELKTIT IKATYREDII RFRLPLDSGI DKLKEEIAKR 960 LKLDVGTFEI KYLDDDHEWV LIACDADLQE CVDISKSSGS NMIRLLVHDL MPALGSSCES 1020 SGECGGLGCK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
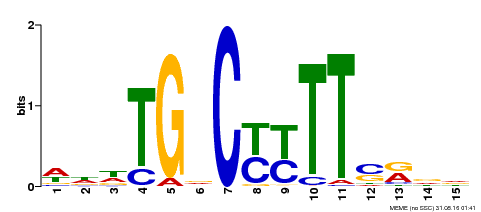
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027085451.1 | 0.0 | protein NLP6-like | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A068UUL3 | 0.0 | A0A068UUL3_COFCA; Uncharacterized protein | ||||
| STRING | XP_009605047.1 | 0.0 | (Nicotiana tomentosiformis) | ||||



