 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_7205_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 324aa MW: 36129 Da PI: 6.7733 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.5 | 1.3e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd lv +v+++G+g+W++++ g++R+ k+c++rw +yl
cra_locus_7205_iso_5_len_1476_ver_3 14 KGPWTPEEDIMLVSYVQEYGPGNWRAVPTNTGLRRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.6 | 1.9e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T +E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
cra_locus_7205_iso_5_len_1476_ver_3 67 RGSFTDQEEKMIIQLQALLGNK-WAAIASYLP-ERTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.0E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 25.751 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.4E-31 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.4E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.0E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.76E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.3E-25 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.3E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.289 | 66 | 116 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.1E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.31E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
MGRPPCCDKV GVKKGPWTPE EDIMLVSYVQ EYGPGNWRAV PTNTGLRRCS KSCRLRWTNY 60 LRPGIKRGSF TDQEEKMIIQ LQALLGNKWA AIASYLPERT DNDIKNYWNT HLKKKLKMLQ 120 TGPDPCSRNG FSSSQSASKG QWERRLQADI KTAKQALQDA LTLDGTSSPV PDSKGLFSLA 180 ETGGQASNNS NTYASSTENI ARLLKDWARN SSNSKSKKQS KSSSTQHSFN YPANEYSTSS 240 SVESKNGIEL SEAFESLFGF ESFDSSNSEF SQSTSPEASL FQGESKPEIT QVPLAMLENW 300 LFDESSIQGK DLTNFSFDHD EDLF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h88_C | 2e-27 | 14 | 116 | 58 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 2e-27 | 14 | 116 | 58 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h8a_C | 6e-28 | 14 | 116 | 27 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 117 | LKKKLKM |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
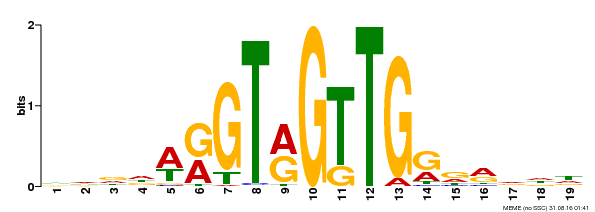
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028109092.1 | 1e-147 | myb-related protein 306-like | ||||
| Swissprot | P81392 | 1e-115 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | A0A2N9FVI2 | 1e-145 | A0A2N9FVI2_FAGSY; Uncharacterized protein | ||||
| STRING | XP_009629125.1 | 1e-139 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



