 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_6055_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 429aa MW: 46431.6 Da PI: 7.1664 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 126 | 1.2e-39 | 71 | 131 | 2 | 62 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkkss 62
++k+l+cprC+s++tkfCyynny+++qPr+fCk+C+ryWt+GG++rnvPvG+grrknk+s
cra_locus_6055_iso_2_len_1645_ver_3 71 PDKILPCPRCNSMDTKFCYYNNYNVNQPRHFCKSCQRYWTAGGTMRNVPVGAGRRKNKNSA 131
7899******************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 8.0E-30 | 69 | 128 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 2.5E-32 | 73 | 129 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.919 | 75 | 129 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 77 | 113 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 429 aa Download sequence Send to blast |
XCLAGDCETK PEEKDEHSIS EETENSKPLS ASDDSPDALP MDEDSPTKES GNAESENEQN 60 DGTNSQQKKK PDKILPCPRC NSMDTKFCYY NNYNVNQPRH FCKSCQRYWT AGGTMRNVPV 120 GAGRRKNKNS ASHCRHITIS EALQAARIDA PNGFHHPTLK PNGTVLSFGP DIPLCDSMAS 180 VLNLAEKKLP NGDRNGFYKV DPVISVSYKG TENGDDCSSG SSVTTSNSMV ENGKKGQQEP 240 VIQHINGFHS PVPCLPGVPW PFPWNSGVPL PAICPTAFPM PFYPAPFPWN CNVASVPGAW 300 SIPWMPPVSP TASQKPSGSS PNSPLGKHSR DGELLNKPNS TKANESPEKK NSESSILVPK 360 TLRIDDPDEA AKSSIWTTLG IKYDSISRGG LFKALQPKGD QKKPLPATSP VLQANPAALS 420 RSVTFQETA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
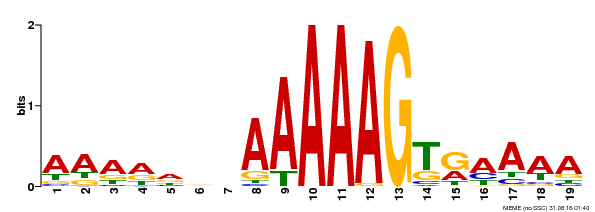
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Regulates a photoperiodic flowering response. Transcriptional repressor of 'CONSTANS' expression. {ECO:0000250, ECO:0000269|PubMed:19619493}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00394 | DAP | Transfer from AT3G47500 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Highly expressed at the beginning of the light period, then decreases, reaching a minimum between 16 and 29 hours after dawn before rising again at the end of the day. {ECO:0000269|PubMed:19619493}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027167719.1 | 0.0 | cyclic dof factor 3 | ||||
| Swissprot | Q8LFV3 | 1e-105 | CDF3_ARATH; Cyclic dof factor 3 | ||||
| TrEMBL | A0A068UVL6 | 0.0 | A0A068UVL6_COFCA; Uncharacterized protein | ||||
| STRING | Solyc03g115940.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



