 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_5882_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 582aa MW: 65977.8 Da PI: 5.8807 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 77 | 2.4e-24 | 200 | 253 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++Wt +LH++Fv+av+q+ G ek Pk+il+lm+v+ Lt+e+v+SHLQkYRl
cra_locus_5882_iso_5_len_2575_ver_3 200 KTRVVWTVDLHQKFVKAVNQI-GFEKVGPKKILDLMGVPWLTRENVASHLQKYRL 253
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 82.4 | 1.5e-27 | 20 | 128 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTS CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepkl 72
vl+vdD+p+ +++l+++l+k y ev+++ ++eal+ll+e++ +D+++ D++mp+mdG++ll++ e +l
cra_locus_5882_iso_5_len_2575_ver_3 20 VLVVDDDPTWLKILEKMLKKCSY-EVTTCGLAREALDLLRERKdgFDIVISDVNMPDMDGFKLLEHVGLEM-DL 91
89*********************.***************999999**********************6644.8* PP
EEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 73 piivvtahgeeedalealkaGakdflsKpfdpeelvk 109
p+i+++ ge + + + ++ Ga d+l Kp+ ++el++
cra_locus_5882_iso_5_len_2575_ver_3 92 PVIMMSVDGETSRVMKGVQHGACDYLLKPIRMKELRN 128
**********************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF52172 | 1.71E-36 | 17 | 151 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 8.3E-45 | 17 | 148 | No hit | No description |
| SMART | SM00448 | 1.3E-32 | 18 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.773 | 19 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.4E-24 | 20 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.01E-30 | 21 | 134 | No hit | No description |
| SuperFamily | SSF46689 | 2.33E-17 | 198 | 257 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-27 | 199 | 258 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-21 | 200 | 253 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.8E-8 | 202 | 252 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 582 aa Download sequence Send to blast |
MEVSGYPSPR TDAFPAGLRV LVVDDDPTWL KILEKMLKKC SYEVTTCGLA REALDLLRER 60 KDGFDIVISD VNMPDMDGFK LLEHVGLEMD LPVIMMSVDG ETSRVMKGVQ HGACDYLLKP 120 IRMKELRNIW QHVFRKRIHE VKEIENHEGF EEIQFMRNVC DQSEDRMFSS ANDLASGKKR 180 KDYENKHDDR ECSDSSSVKK TRVVWTVDLH QKFVKAVNQI GFEKVGPKKI LDLMGVPWLT 240 RENVASHLQK YRLYLTRLQK EKELTASFGV MKHSDISSKE QTGNHCLQNS VNMQQTNGSC 300 GISGNKTAMQ NIDFEIHEGD LKSLVKIPMV DANNGLGGDN PDSQRGRRMN ADHSFRKLDS 360 DVKYATTIPE QYMWSAEGSE VQANQERKQP FRPKPGFNPM PLPSLHQNIQ FDCVRPTPSF 420 VSKPSNIHND KLGGSEKKTI FVKSVSHGNM VDLLPVQPQS ELTATTSQEV LKPDPSSIVW 480 STKNQRAADQ IFVSDMESAQ RSIDLVGGWG LATWEENQQG CLIEADIHRL TQGLQNIESF 540 SYNEAGIISE ASSYLYDSLK FDYEMPLEPM EYSVIDQGLF IV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 4e-17 | 196 | 258 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
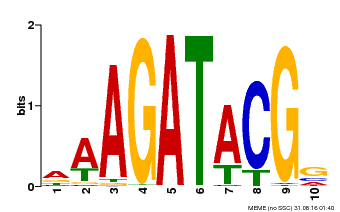
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027100212.1 | 0.0 | two-component response regulator ORR26 isoform X1 | ||||
| Refseq | XP_027152868.1 | 0.0 | two-component response regulator ORR26 | ||||
| TrEMBL | A0A2R6RED2 | 0.0 | A0A2R6RED2_ACTCH; Two-component response regulator like | ||||
| STRING | VIT_01s0010g02230.t01 | 0.0 | (Vitis vinifera) | ||||



