 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_4867_iso_4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 102aa MW: 11701.2 Da PI: 11.1502 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 36.3 | 1.2e-11 | 41 | 86 | 2 | 51 |
XXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 2 kelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkelee 51
++ k rr+++NReAAr+sR+RKka++++Le +L++ ++L ++
cra_locus_4867_iso_4_len_862_ver_3 41 RDQKTLRRLAQNREAARKSRLRKKAYVQQLESSRMKLTQLEQEL----QR 86
567899*************************9999998877644....44 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 2.4E-10 | 37 | 89 | No hit | No description |
| SMART | SM00338 | 1.7E-5 | 40 | 102 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.5E-8 | 42 | 85 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.461 | 42 | 86 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.06E-8 | 44 | 90 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 47 | 62 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0009627 | Biological Process | systemic acquired resistance | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 102 aa Download sequence Send to blast |
MADVSPRTDT STDASTEDKS TRFQNNQGLA IAASDGSDKS RDQKTLRRLA QNREAARKSR 60 LRKKAYVQQL ESSRMKLTQL EQELQRARQQ VCFNKLLMRI LS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
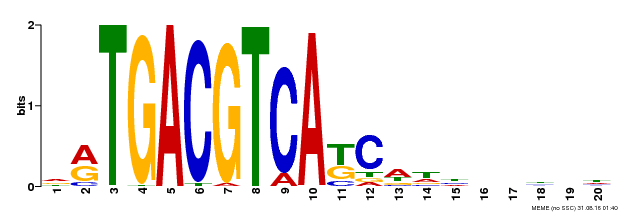
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. Binds to the hexamer motif 5'-ACGTCA-3' of histone gene promoters. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027078869.1 | 2e-52 | transcription factor TGA2.2-like isoform X2 | ||||
| Refseq | XP_027161661.1 | 2e-52 | transcription factor TGA2.2-like | ||||
| Swissprot | Q41558 | 4e-33 | HBP1C_WHEAT; Transcription factor HBP-1b(c1) (Fragment) | ||||
| TrEMBL | A0A068UNV8 | 1e-51 | A0A068UNV8_COFCA; Uncharacterized protein | ||||
| STRING | Migut.F00128.1.p | 3e-45 | (Erythranthe guttata) | ||||



