 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_4298_iso_10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 567aa MW: 62985.4 Da PI: 7.0707 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 23.2 | 1.6e-07 | 536 | 563 | 1 | 28 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIa 28
++rW++ E++ l av+++G g+Wk+I
cra_locus_4298_iso_10_len_1699_ver_3 536 KKRWSALEEDALRSAVEKHGYGNWKLIL 563
68************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-7 | 530 | 563 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 10.623 | 531 | 567 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.26E-7 | 533 | 564 | IPR009057 | Homeodomain-like |
| CDD | cd11660 | 4.62E-6 | 539 | 565 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 567 aa Download sequence Send to blast |
XLSSLKSLLL SFTSFPRKKT PLHVPDADIG DYXRSPHAFP SMSSINTKIF PESRGGSXKA 60 TEGFLYWLSK YSWNKRKHGP RIIADCLLKL LLHIIQSPNL DMDSDLGPWI LESLLRLPLQ 120 SRILKSLIKV VPLPNDNPII KKIILLRKIE SEISKGCVSE SILDCLERIE ELDFQSGMDH 180 DSESLKAAYC AVAVTCTVQS IADKGADSKF KYFEAVKRIW KEKVCKMEKV ENVGLISEEL 240 LSWKDEIEAA LWEESVCDDV LKRTEGMDAV GAVGAYVHEE KGKMGPSFLE FVAQKARTDD 300 TLMKLLFDVG NANLGQGGEK KENLAIAADQ GLGRSDNREG HKEIVLSKAK HIALKRTRGP 360 RFGISRGSKI ADTGNSEVDP SGKHCDLLPT PEVNKVQEDL RSSSLELQAV VKDPLPDALN 420 LAEMLLSFTE KENTSKEIVV LSKVGDDPSF VESSKAVQAN EEDLSKHCQD VRTNAAETSL 480 MESNGTVRAF ECDDSIDETP DISPSGKGLR LPSPKGVNVS PLKKYEANKL TKRRKKKRWS 540 ALEEDALRSA VEKHGYGNWK LILMTNR |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 531 | 537 | KRRKKKR |
| 2 | 532 | 537 | RRKKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
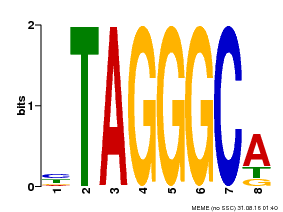
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027091291.1 | 1e-159 | uncharacterized protein LOC113712180 isoform X2 | ||||
| Refseq | XP_027112461.1 | 1e-159 | uncharacterized protein LOC113731391 isoform X2 | ||||
| TrEMBL | A0A068U6V9 | 1e-156 | A0A068U6V9_COFCA; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3774 | 24 | 39 |



