 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_3631_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 338aa MW: 35925.6 Da PI: 10.3605 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 72.4 | 6.5e-23 | 195 | 257 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkrerrkq+NRe+ArrsR+RK+ae+eeL++kv++L+aeN++L++e+++l+++++ l+ e+
cra_locus_3631_iso_2_len_1684_ver_3 195 ERELKRERRKQSNRESARRSRLRKQAETEELAKKVQTLTAENMTLRSEINKLTENSEHLRHEN 257
89********************************************************99887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF16596 | 3.1E-25 | 43 | 176 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 4.2E-17 | 190 | 255 | No hit | No description |
| SMART | SM00338 | 8.6E-22 | 195 | 259 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 3.8E-21 | 195 | 257 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.472 | 197 | 260 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.06E-11 | 198 | 254 | No hit | No description |
| CDD | cd14702 | 1.81E-18 | 200 | 248 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 202 | 217 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 338 aa Download sequence Send to blast |
MGSSEETKSS KPEKSSSPGA TEAIAASPLS IDTPTKSSAN GSQGLMNKLR GFDGLAMSIG 60 NGNTDSADGG TDHGISQSGD TEGSSDGSNG TTSKAGQKNK KRSREGTPAN DRERKSLTPS 120 SPSAAVNTNG SSEKAMRASK VPAAATEKVM GAVLSPNMTT ASELRNPSAA NAKTSPAKVP 180 NPVLLFRVKA WLQNERELKR ERRKQSNRES ARRSRLRKQA ETEELAKKVQ TLTAENMTLR 240 SEINKLTENS EHLRHENAAL LDKLKNARVM QAGEMNKYDE LHRQPTGTAD LLARVNNSGS 300 TDKSNEEGGG DVFENRNSGT KLHQLLDASP RADAVAAG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 211 | 217 | RRSRLRK |
| 2 | 211 | 218 | RRSRLRKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
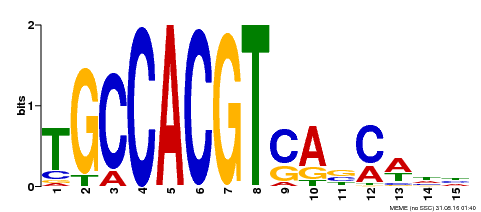
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027108496.1 | 1e-138 | transcriptional activator TAF-1-like isoform X3 | ||||
| TrEMBL | Q9XHQ6 | 0.0 | Q9XHQ6_CATRO; G-box binding protein 1 | ||||
| STRING | VIT_15s0046g01440.t01 | 1e-122 | (Vitis vinifera) | ||||



