 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_2886_iso_4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 464aa MW: 49089.6 Da PI: 9.9736 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 48.2 | 2.4e-15 | 377 | 428 | 5 | 56 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkev 56
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++eN++L+k+ e+ ++
cra_locus_2886_iso_4_len_1870_ver_3 377 RRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKEENQELRKKQAEIIEMQ 428
79****************************************9987776665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.4E-14 | 373 | 439 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.461 | 375 | 424 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 4.1E-15 | 377 | 424 | No hit | No description |
| CDD | cd14707 | 1.50E-26 | 377 | 431 | No hit | No description |
| SuperFamily | SSF57959 | 8.13E-11 | 377 | 425 | No hit | No description |
| Pfam | PF00170 | 5.1E-13 | 377 | 429 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 380 | 395 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 464 aa Download sequence Send to blast |
MDGGGGGSAG GRPPGNYPLT RQPSVYSLTF EEFQNTIGGS GKDFGSMNMD ELLKNIWSAE 60 EIQSIGSATT AVQDGGASGG YLQRQGSLTL PRTLSQKTVD EVWKDISKEF IGTAGGGKDG 120 SAVGGSSVAQ TQRQQTLGEV TLEEFLVRAG VVREEAQLAA RPNTAGLFAD FSRTGNNQGL 180 GFANQQPGRN TGYMGGRILE SSNQIAAESA NLPLNXKWGK IYPAEVRDTT NTAASTTPGA 240 AAPAPAQHQH QHQQQPIFPK QPTLAYGAPM GIPNGAQMAI PNGAQLGSPS LRGGIVGISD 300 PITNGNLVQN AALQGGGMGM VGLGAGGVTV AAGSPAVSSD GLGKSNGDAS SVSPVPYVFN 360 GGLRGRKCSA LEKVVERRQR RMIKNRESAA RSRARKQAYT MELEAEVAKL KEENQELRKK 420 QAEIIEMQKN QAMEMMNMQL GGKRRCLRRT QTGPWXREHR MLEP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
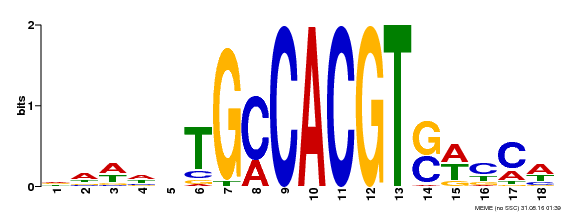
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027091459.1 | 0.0 | bZIP transcription factor 46-like | ||||
| Refseq | XP_027091460.1 | 0.0 | bZIP transcription factor 46-like | ||||
| Swissprot | Q9M7Q4 | 1e-127 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | A0A1U8GBR9 | 0.0 | A0A1U8GBR9_CAPAN; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | A0A1U8GJ68 | 0.0 | A0A1U8GJ68_CAPAN; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| STRING | Solyc04g078840.2.1 | 1e-179 | (Solanum lycopersicum) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



