 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_2781_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1043aa MW: 116620 Da PI: 6.7002 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.5 | 9.8e-57 | 21 | 136 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvg 76
+ ++rwl++ ei++iL n++k+++t e++ +p sgs++L++rk++ryfrkDG+sw+kkkdgktv+E+hekLKvg
cra_locus_2781_iso_5_len_3547_ver_3 21 EAQHRWLRPAEICEILRNYQKFHITPEPPLKPVSGSVFLFDRKVLRYFRKDGHSWRKKKDGKTVKEAHEKLKVG 94
569*********************************************************************** PP
CG-1 77 gvevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
++++l+cyYah+een++fqrr+yw+Le++l +iv+vhylevk
cra_locus_2781_iso_5_len_3547_ver_3 95 SIDMLHCYYAHGEENENFQRRSYWMLEQDLMHIVFVHYLEVK 136
***************************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.983 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.5E-79 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.1E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.0E-7 | 483 | 571 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.8E-9 | 484 | 570 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.52E-18 | 485 | 571 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 7.0E-17 | 666 | 775 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.13E-17 | 670 | 775 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.58E-15 | 674 | 772 | No hit | No description |
| PROSITE profile | PS50297 | 17.369 | 680 | 774 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 1.2E-6 | 685 | 775 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.671 | 713 | 739 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0032 | 713 | 742 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1500 | 752 | 781 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.47E-7 | 887 | 939 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.17 | 888 | 910 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.84 | 889 | 918 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0025 | 890 | 909 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.003 | 911 | 933 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.432 | 912 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.9E-4 | 914 | 933 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1043 aa Download sequence Send to blast |
MAGSGSQNLG FRLDIKQILC EAQHRWLRPA EICEILRNYQ KFHITPEPPL KPVSGSVFLF 60 DRKVLRYFRK DGHSWRKKKD GKTVKEAHEK LKVGSIDMLH CYYAHGEENE NFQRRSYWML 120 EQDLMHIVFV HYLEVKGNKT NINCLLDSET VSSNSQNDTL SFSSSANGDN LAAAYTDTTS 180 PTSTLTSAYE ETECAQQNVE DNYRPSSRVH SHPGTPQSES IPAAQHVDAG LSSSYSLLQS 240 LGTQMTPSGE YIPFSHGQGG AHGHEGSFVP GIQRTLDLAS WEEVWGNSTT AEISSDHHSW 300 NPSGLQANWQ YPQGDSSLSF QGLPEQNLIS SSASDDRGNF LDHRSIPANF QRLADPFYVN 360 AEGLEKELVQ REHQKLLSNA QISYMKKDKA ENGNAAMGSG SLVLKQSQLS GIKIDDGLKK 420 VDSFSRWMAK ELGEVEELHL QTNNGYSWSE IQTEDVVDSS GMPSQLQLEA DTLNFSLSQD 480 QLFSITDFSP NWAYSNSETK VIITGRYLKS DQLFVNCKWS CMFGEVEVPA VVLGEGVLCC 540 HAPRHKPGLV PFYVTCSNRL ACSEVRDFEY RAGPQDIPRG DTDVMHICRR LDRLLSIGPV 600 VSINNFSENV VERNDIVKKI ISLMEEDDNQ MPKLASGKDI SLLKAVEVQQ AEKWLRDKFY 660 EWILNKVKED GRGPSIIDDE GQGVLHLAAA LGFNWALEPI IVSGVSIDFR DVNGWTALHW 720 AAFYGREDTV AALVSLGASC GPLTDPTAEH PLGRTPADLA SANGHKGISG FLAECSLTAH 780 LELLTVKETE EDSTLEYSGA NAIQTVSERV AAPISGGDVP DSLSLKDSLA AVCNATQAAA 840 RIHQIFRVQS FQRKQLIERT VNDDASSDEY ALSILAAKTS RLGQNDVTAH AAAVSIQKKY 900 RGWKKRKEFL IIRQRIVKIQ AHVRGHQVRK KYKPIIWSVG ILEKVILRWR RKGSGLRGYR 960 PDAVAKVPGE LDMPQKEDDY DFLKEGRKQT EERLQKALTR VKSMAQYPEA RAQYRRLLTV 1020 AEGLRQTKEN NYLLKSRFVY KVN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 2e-13 | 485 | 574 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_B | 2e-13 | 485 | 574 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_C | 2e-13 | 485 | 574 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_D | 2e-13 | 485 | 574 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_E | 2e-13 | 485 | 574 | 9 | 92 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
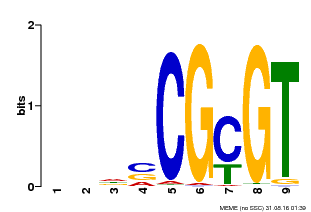
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027106275.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A068TQ04 | 0.0 | A0A068TQ04_COFCA; Uncharacterized protein | ||||
| STRING | XP_009600617.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



