 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_23355_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 401aa MW: 44074.2 Da PI: 7.7908 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.7 | 4.8e-32 | 293 | 348 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+p+LH+rFv+a++ LGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
cra_locus_23355_iso_1_len_1202_ver_3 293 KARRCWSPDLHRRFVNALQMLGGSQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 348
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.204 | 290 | 350 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.73E-17 | 291 | 351 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-28 | 291 | 351 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.0E-26 | 293 | 348 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.6E-7 | 295 | 346 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 401 aa Download sequence Send to blast |
EEKLKLMMAS PSELSLECKT QYSCSMLLKS VGGEQSDIQT QKLEDFLARL EEERLKIDAF 60 KRELPLCMQL LTNAMEASRQ QLHSYRANNN EGPKPILEEF IPLKHHNPAI SEGSEKTNNN 120 NTNVNVVSDK ANWMTSAQLW SQASSGGTSA GGVDHEIIAK QQQQSPAAAV TSPKESDDHM 180 GFGVNLKQSE NNNNNNNQRN GNGNGGGGFL PFSKERNLNR GGAVVVPELA LSSTDHHHNH 240 HEIMEEKKII NSEGTSSDNM MMMMRRDQNL LGNNNNKSAA GESQTTTNQS HRKARRCWSP 300 DLHRRFVNAL QMLGGSQVAT PKQIRELMKV DGLTNDEVKS HLQKYRLHTR RPSPSPQSAA 360 AAAPQLVVLG GIWVPPEYAT AAAAHHAGGP PTLYPTTHSS P |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 8e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-14 | 293 | 346 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a flowering repressor, directly repressing FT expression in a dosage-dependent manner in the leaf vasculature. {ECO:0000269|PubMed:25132385}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
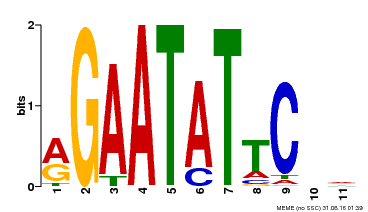
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by SVP. Down-regulated by high temperature and gibberellic acid treatment. Not regulated by photoperiod, circadian rhythm under long days or vernalization. {ECO:0000269|PubMed:25132385}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017422950.1 | 1e-137 | PREDICTED: myb family transcription factor EFM | ||||
| Swissprot | Q9ZQ85 | 1e-112 | EFM_ARATH; Myb family transcription factor EFM | ||||
| TrEMBL | A0A218W107 | 1e-136 | A0A218W107_PUNGR; Uncharacterized protein | ||||
| STRING | XP_004508555.1 | 1e-133 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8794 | 22 | 29 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



