 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_23047_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 349aa MW: 39024 Da PI: 6.9085 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 83.4 | 1.8e-26 | 102 | 167 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kfl+l+++++ g+++ln+ a+ L +v ++RRiYDi+NVLe+++liek+ kn+irw+g
cra_locus_23047_iso_1_len_1142_ver_3 102 RYDSSLGLLTKKFLNLIQEAPGGTLDLNKTADVL---EV--QKRRIYDITNVLEGIGLIEKTTKNHIRWRG 167
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 3.8E-28 | 99 | 168 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 1.1E-35 | 102 | 167 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF46785 | 7.71E-18 | 102 | 165 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF02319 | 6.7E-25 | 104 | 167 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF144074 | 8.63E-26 | 180 | 286 | No hit | No description |
| CDD | cd14660 | 2.43E-36 | 183 | 285 | No hit | No description |
| Pfam | PF16421 | 1.7E-24 | 183 | 283 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000902 | Biological Process | cell morphogenesis | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042023 | Biological Process | DNA endoreduplication | ||||
| GO:0051782 | Biological Process | negative regulation of cell division | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MSNTHEINLN LTLGTLNTSF SNVSLESQQS RCSVPQLLAS CSSRRSPFLP SASSTTAAFP 60 DSKHALATAS PPKARGSKLA KSGTVEANAE ILENLGLSGS CRYDSSLGLL TKKFLNLIQE 120 APGGTLDLNK TADVLEVQKR RIYDITNVLE GIGLIEKTTK NHIRWRGSEL LGPRELDHQA 180 NFLKAEVEHL YAEECRLDNR IREKLEQLRA LEFDQTSQRN LFLTKEDIMN LPCFRDKTVI 240 AVKAPHASNI EVPDPEEEMD LSGKKHYRLI LRSNTGPIDL YLLSKNQKKH EDLAAKRRKS 300 SDLWVSSVSR HRIDNAEASG IHKIIPLDDA DGDDYWLRSD HEATATDLW |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 2e-22 | 102 | 173 | 7 | 76 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in transcriptional repression. May act by repressing E2F-regulated genes in mature differentiated cells, but is not an antagonist of E2FA. Restricts cell division and is involved in the coordination between cell proliferation and endoreduplication during development. May play a role during the transition from skotomorphogenesis to photomorphogenesis. Regulated by phosphorylation-dependent proteolysis via the protein-ubiquitin ligase SCF(SKP2A) complex. {ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12468727, ECO:0000269|PubMed:16920782, ECO:0000269|PubMed:19662336}. | |||||
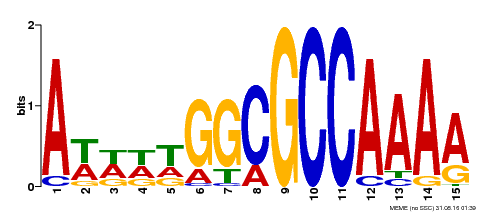
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00190 | DAP | Transfer from AT1G47870 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by light. {ECO:0000269|PubMed:12468727, ECO:0000269|PubMed:18424613}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027092083.1 | 1e-122 | transcription factor E2FC-like | ||||
| Refseq | XP_027094734.1 | 1e-122 | transcription factor E2FC-like | ||||
| Swissprot | Q9FV70 | 4e-87 | E2FC_ARATH; Transcription factor E2FC | ||||
| TrEMBL | A0A2G9HCT7 | 1e-113 | A0A2G9HCT7_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.C00805.1.p | 1e-114 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12625 | 20 | 22 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



