 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_22402_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 299aa MW: 33869.5 Da PI: 6.1794 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 57.7 | 2.6e-18 | 15 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWT+eEde+l++++kq G g+W++ ++ g+ R++k+c++rw +yl
cra_locus_22402_iso_1_len_896_ver_3 15 RGRWTAEEDEILINYIKQNGEGSWRSLPKNAGLLRCGKSCRLRWINYL 62
8*******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.2 | 1.6e-17 | 68 | 113 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+++ eE+e ++++++ G++ W++Ia++++ gRt++++k++w+++l
cra_locus_22402_iso_1_len_896_ver_3 68 RGNFSVEEEETIIKLHTSVGNR-WSLIASHLP-GRTDNEVKNYWNSHL 113
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-23 | 6 | 65 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.866 | 10 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.6E-29 | 12 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.7E-13 | 14 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-15 | 15 | 62 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.86E-10 | 17 | 62 | No hit | No description |
| PROSITE profile | PS51294 | 25.873 | 63 | 117 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-25 | 66 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.6E-16 | 67 | 115 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.5E-15 | 68 | 113 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.44E-11 | 70 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0051555 | Biological Process | flavonol biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
KMGRAPCCEK VGLKRGRWTA EEDEILINYI KQNGEGSWRS LPKNAGLLRC GKSCRLRWIN 60 YLRADLKRGN FSVEEEETII KLHTSVGNRW SLIASHLPGR TDNEVKNYWN SHLSRKIYSF 120 KTNNGSLLNS RDVLQMTKST TSKRRGGRVS RAIAKKYNKP IPLEIQQSQE THDSCYEGQE 180 VLFHDSILMG SSSKELGGGG GEIEVMGTYE DDHQTSTDDV TTLVDQFFLS ELSDLYKISI 240 NNDEQKVNAT TTSMVPENSI NSEEWKSETW HSISNSSLDS GLLDECFSPL NSYFDDMVX |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 6e-24 | 13 | 117 | 5 | 108 | B-MYB |
| 1h8a_C | 9e-24 | 13 | 117 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Flavonol-specific transcription activator involved in the regulation of several genes of flavonoid biosynthesis. Activates the expression of CHS, CHI, F3H and FLS1. Controls flavonol biosynthesis primarily in cotyledons and leaves (PubMed:17419845, PubMed:20731781). Confers tolerance to UV-B (PubMed:19895401). {ECO:0000269|PubMed:17419845, ECO:0000269|PubMed:19895401, ECO:0000269|PubMed:20731781}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
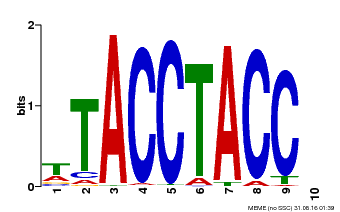
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Triggered by HY5 in response to light and UV-B. {ECO:0000269|PubMed:19895401}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027119798.1 | 5e-89 | myb-related protein 330-like isoform X1 | ||||
| Refseq | XP_027164906.1 | 5e-89 | myb-related protein 330-like | ||||
| Swissprot | Q9FJ07 | 1e-73 | MY111_ARATH; Transcription factor MYB111 | ||||
| TrEMBL | A0A068TR64 | 1e-87 | A0A068TR64_COFCA; Uncharacterized protein | ||||
| STRING | Migut.N00057.1.p | 8e-81 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |



