 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_18283_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 894aa MW: 102231 Da PI: 7.4953 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.4 | 5e-28 | 137 | 224 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekd 73
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e++ + ++r+ ++++t+Cka+++vk++kd
cra_locus_18283_iso_5_len_3321_ver_3 137 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESDGT---SSRRASVKKTDCKASMHVKRKKD 206
5****************************************999888...8889999**************** PP
FAR1 74 gkwevtkleleHnHelap 91
gkw+v+++ +eHnHel p
cra_locus_18283_iso_5_len_3321_ver_3 207 GKWYVHEFIKEHNHELLP 224
***************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 8.8E-26 | 137 | 224 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 5.8E-26 | 342 | 435 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.99 | 622 | 658 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.9E-7 | 624 | 657 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 8.3E-9 | 633 | 660 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 894 aa Download sequence Send to blast |
XNLLTTDXSG RGPILHPPLS TCVAADHFYA QTVVICSATV DLRAKRNCFC DILQPGHFVF 60 CRKGISEGSS EMHLGAWCGE EVXIAVVMIG NMVDVVDHAR DRNARHSPQR DMVGTSGDTE 120 FEPHDGIEFE SHEAAYSFYQ EYAKSMGFTT SIKNSRRSKK SKEFIDAKFA CSRYGVTPES 180 DGTSSRRASV KKTDCKASMH VKRKKDGKWY VHEFIKEHNH ELLPALAYHF RIHRNVKLAE 240 KNNIDILHAV SERTRKMYVE MSRQSGGNPD IDFLRNSFKH EGRQCLTLEE GDAQVMLEYF 300 MHVQRENPYF FYAMDLNEDQ RLRNLFWVDA KSRKDYNSFS DVILFDTSYI KSNEKMPFAP 360 FVGVNNHSQP MLLGCALIAD ESTSTIVWLM KTWLRAMGGQ PPKVIITDQG KTLRAAIAEI 420 FPQSRHCYAL WHILEKVPEI LAHVIKQHET FLGKLHKCIF KSLNDEDFDM RWWKMIGRFE 480 LQENEWVHSL YEDRKKWVPA FTKDAFLAGL STNQRSDSVT SFFDKYIHKK ISLKEFVRQY 540 GTILQNRYEE EAIADFDTWH KQPALKSPSP WEKQMSAIYT HAIFKKFQVE VLGVVGCHPK 600 KESVSGTDIS FRVDDCEKGE NFIVTWNEEA LNVSCSCLMF ESKGFLCRHA LIVLQMCGLS 660 SIPSQYILKR WTKDAKNRPT ILDGTERNQT RVQRYNDICR RAIELGEEGS LSEESYNIAF 720 RALVEGLKNC VDVNNKSAVE CSSNPVGLRD VEEDNQDIHS KSSKKKSTNK KRRVQPDSEA 780 AIVEVQGSLH PMENLNSDTM TLNGYYGTQQ NVPGLIQLNL MEPPREGYYV NQQNMQGLGQ 840 LSSIAPSHDG FYATPQSIPA LGHLDFRPPS FTYSLQVVKF IGISYFNSLS LITF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
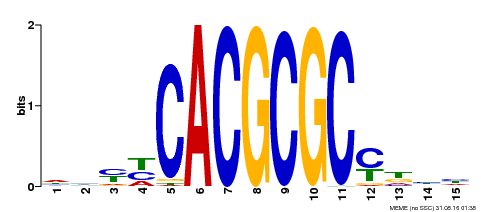
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027171325.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X2 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A1S4C3S0 | 0.0 | A0A1S4C3S0_TOBAC; protein FAR-RED IMPAIRED RESPONSE 1-like isoform X1 | ||||
| TrEMBL | A0A1U7VQ31 | 0.0 | A0A1U7VQ31_NICSY; protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| STRING | XP_009767065.1 | 0.0 | (Nicotiana sylvestris) | ||||



