 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_17221_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 359aa MW: 41168.9 Da PI: 7.7721 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.6 | 5.2e-32 | 70 | 123 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+++AtPk +l+lm++kgL ++hvkSHLQ+YR+
cra_locus_17221_iso_2_len_1399_ver_3 70 PRLRWTPDLHLRFVHAVERLGGQDRATPKLVLQLMNIKGLNIAHVKSHLQMYRS 123
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.354 | 66 | 126 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-30 | 66 | 124 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.85E-15 | 68 | 124 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.2E-22 | 70 | 124 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.3E-8 | 71 | 122 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 359 aa Download sequence Send to blast |
MEGSGDTTEC SKNSKNEDED ECYEEEDEKF DEQKQKKLDG GSSSNSTVDE NNNHHDKKPS 60 VRPYVRSKMP RLRWTPDLHL RFVHAVERLG GQDRATPKLV LQLMNIKGLN IAHVKSHLQM 120 YRSKRIDDPS LGIGDHRHLF EGAADRNIYN LSQLPMLRGF NQNHSSPPFR YNRDACWNGL 180 QNLMHNTSMG QSTINKMRIL GSNYSRIMDS SQGSSWKKDE LKSFLDKEAW KGNYNHMDQL 240 KTLKHQEKKS RIDHIQDLMI TSSSNTNSSR LEKEGVRKRK AITESDEIDL TLSLRLGTKT 300 EDEDSFGSQL EDSDNESISY LSLSLFSPSS SSKKIRRRMK EDEIIRENGK GASTLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 2e-17 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-17 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-17 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-17 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
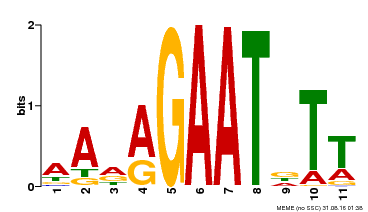
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011095220.1 | 4e-91 | uncharacterized protein LOC105174731 | ||||
| TrEMBL | A0A068UFP0 | 5e-89 | A0A068UFP0_COFCA; Uncharacterized protein | ||||
| STRING | EOY07117 | 5e-80 | (Theobroma cacao) | ||||



