 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_15774_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 279aa MW: 30287.8 Da PI: 9.2243 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 105 | 4.3e-33 | 36 | 90 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LHerFv+av++LGG++kAtPk++l++m++kgLtl+h+kSHLQkYRl
cra_locus_15774_iso_2_len_1400_ver_3 36 KPRLRWTADLHERFVDAVTKLGGPDKATPKSVLRVMGLKGLTLYHLKSHLQKYRL 90
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.0E-32 | 33 | 91 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.656 | 33 | 93 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.8E-24 | 36 | 91 | IPR006447 | Myb domain, plants |
| SuperFamily | SSF46689 | 3.94E-16 | 36 | 93 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.3E-9 | 38 | 89 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.7E-24 | 136 | 182 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MESMFGGGGG GMGGGCYPYE RNGNGGAAVV MTRDPKPRLR WTADLHERFV DAVTKLGGPD 60 KATPKSVLRV MGLKGLTLYH LKSHLQKYRL GQQAKKQNGA EQNKENSGDS FGQFSSQASA 120 SSSHSTRMNI EPGEIPITEA LKSQIEVQKR LQEQLEVQKK LQMRIEAQGK YLQTILEKAQ 180 KSLSVEANNG NNVEATRAEL SEFNLALSSL IENVKGEERK GGNVVDSRRK ICSSSSSYYG 240 AKDDDSNDVK LTVEGGGSIN FDLNTRSSSY DFIGNSGAV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 2e-19 | 36 | 89 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-19 | 36 | 89 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-19 | 36 | 89 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-19 | 36 | 89 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
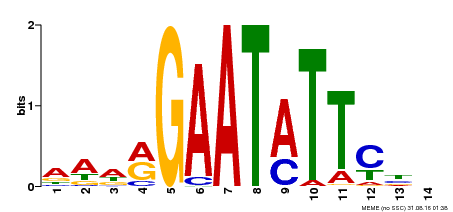
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027084214.1 | 1e-128 | myb family transcription factor PHL11-like | ||||
| Swissprot | C0SVS4 | 6e-81 | PHLB_ARATH; Myb family transcription factor PHL11 | ||||
| TrEMBL | A0A068UVY1 | 1e-125 | A0A068UVY1_COFCA; Uncharacterized protein | ||||
| STRING | EOY25282 | 1e-103 | (Theobroma cacao) | ||||
| STRING | Solyc08g076010.2.1 | 1e-103 | (Solanum lycopersicum) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



