 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_15316_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 259aa MW: 29168.7 Da PI: 10.175 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.3 | 6.2e-17 | 13 | 60 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd ll ++v+++G g W+ ++ + g++R++k+c++rw++yl
cra_locus_15316_iso_1_len_800_ver_3 13 KGAWTEEEDILLRKCVEKYGEGKWHQVPFKAGLNRCRKSCRLRWLNYL 60
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.9 | 1.8e-16 | 66 | 111 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++ +Ed+l+++++k+lG++ W++Ia +++ gRt++++k++w+++l
cra_locus_15316_iso_1_len_800_ver_3 66 RGKFNSDEDDLILRLHKLLGNR-WSLIAGRLP-GRTANDIKNYWNSHL 111
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 25.697 | 8 | 64 | IPR017930 | Myb domain |
| SMART | SM00717 | 6.0E-14 | 12 | 62 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 8.8E-30 | 12 | 107 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.9E-16 | 13 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-23 | 14 | 67 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.38E-11 | 15 | 60 | No hit | No description |
| SMART | SM00717 | 5.3E-15 | 65 | 113 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.417 | 65 | 115 | IPR017930 | Myb domain |
| Pfam | PF00249 | 6.0E-15 | 66 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.29E-11 | 68 | 111 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-25 | 68 | 115 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 259 aa Download sequence Send to blast |
XGSSGSAAVV LRKGAWTEEE DILLRKCVEK YGEGKWHQVP FKAGLNRCRK SCRLRWLNYL 60 RPNINRGKFN SDEDDLILRL HKLLGNRWSL IAGRLPGRTA NDIKNYWNSH LQKKLNGSSK 120 NKNKKNNKSQ KIKKAMITNN NIIKPQPRTF SKNAPWLRAK QVIHNPSPAA AQDSAAPKSQ 180 SMAAQDSADM MTWLDNLLND EQIISNDQGA IWSFNGGSSS EINPGRDQNE DILINQENDL 240 LDFCIDADIW AISSSPANI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 4e-24 | 10 | 115 | 1 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 4e-24 | 10 | 115 | 1 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH002/EGL3/MYC146, BHLH012/MYC1, or BHLH042/TT8. {ECO:0000269|PubMed:15361138}. | |||||
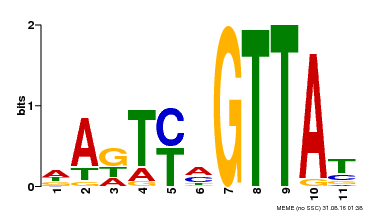
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00213 | DAP | Transfer from AT1G66370 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002274992.2 | 9e-76 | PREDICTED: transcription factor MYB90 | ||||
| Swissprot | Q9FNV9 | 3e-63 | MY113_ARATH; Transcription factor MYB113 | ||||
| TrEMBL | B8YPF8 | 2e-74 | B8YPF8_VITVI; R2R3 MYBA6 transcription factor splice variant 1 | ||||
| STRING | VIT_14s0006g01290.t01 | 3e-75 | (Vitis vinifera) | ||||
| STRING | Lus10028513 | 2e-74 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



