 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_14567_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 204aa MW: 22863.8 Da PI: 8.841 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 63.3 | 4.6e-20 | 25 | 72 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +l+++++ +G++ Wkt+a + g++R++k+c++rw++yl
cra_locus_14567_iso_2_len_901_ver_3 25 KGAWTAEEDRRLAQYIEIHGPKKWKTVAMKAGLNRCGKSCRLRWLNYL 72
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.3 | 5.6e-16 | 78 | 123 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ + eE++l+++++k+lG++ W++Ia +++ gRt++++k++w+++l
cra_locus_14567_iso_2_len_901_ver_3 78 RGNISIEEEDLILRLHKLLGNR-WSLIAGRLP-GRTDNEIKNYWNSHL 123
678889****************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.166 | 20 | 72 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.77E-30 | 24 | 119 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.1E-16 | 24 | 74 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.9E-18 | 25 | 72 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-24 | 26 | 79 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.73E-12 | 27 | 72 | No hit | No description |
| PROSITE profile | PS51294 | 25.738 | 73 | 127 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.9E-14 | 77 | 125 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.2E-14 | 78 | 123 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-25 | 80 | 127 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.50E-11 | 84 | 123 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009957 | Biological Process | epidermal cell fate specification | ||||
| GO:0010053 | Biological Process | root epidermal cell differentiation | ||||
| GO:0010091 | Biological Process | trichome branching | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0048629 | Biological Process | trichome patterning | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 204 aa Download sequence Send to blast |
MAPIKLSAAA EGCSSNPKRT VTNNKGAWTA EEDRRLAQYI EIHGPKKWKT VAMKAGLNRC 60 GKSCRLRWLN YLRPNIKRGN ISIEEEDLIL RLHKLLGNRW SLIAGRLPGR TDNEIKNYWN 120 SHLSKKVNST KRSDVVSASI QESPPQTTTN STSSAEAEPQ PTQLGHGDDM ADLSFDVNDF 180 FDFSVEGTFG MDWISKFLEL DGNP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 2e-30 | 24 | 127 | 26 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH2/EGL3/MYC146 or BHLH12/MYC1. Regulates the epidermal cell fate specification. Mediates the formation of columellae and accumulation of mucilages on seed coats. Controls the elongation of epidermal cells positively in roots but negatively in stems, leading to the promotion of primary roots elongation and repression of leaves and stems elongation, respectively. Ovoids ectopic root-hair formation, probably by inducing GL2 in roots. Controls trichome initiation and branching. {ECO:0000269|PubMed:11437443, ECO:0000269|PubMed:15361138, ECO:0000269|PubMed:15668208, ECO:0000269|PubMed:15728674}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00538 | DAP | Transfer from AT5G40330 | Download |
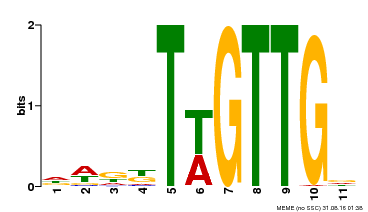
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024169376.1 | 7e-88 | transcription factor WER-like | ||||
| Swissprot | Q96276 | 4e-53 | MYB23_ARATH; Transcription factor MYB23 | ||||
| TrEMBL | A0A2U9IY24 | 1e-150 | A0A2U9IY24_CATRO; MYB domain containing protein transparent testa 2a | ||||
| STRING | XP_008364573.1 | 5e-86 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



