 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_14522_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 243aa MW: 27180.6 Da PI: 9.3417 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 122.5 | 5.1e-38 | 135 | 220 | 4 | 90 |
DUF702 4 gtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasaaeaaskrkr 76
++sCqdCGnqakkdCah+RCRtCCksrg++C thvkstWvpaakrrerqqqla+++++++++++++++++++
cra_locus_14522_iso_2_len_1609_ver_3 135 SGISCQDCGNQAKKDCAHMRCRTCCKSRGYNCPTHVKSTWVPAAKRRERQQQLASLQQQQQHQQNQQQQQQHQ 207
5689****************************************************99998887777666666 PP
DUF702 77 elkskkqsalsstk 90
+ ++++ ++tk
cra_locus_14522_iso_2_len_1609_ver_3 208 LQLHHHHH-RENTK 220
55333333.33332 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 4.9E-35 | 136 | 227 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 6.5E-28 | 138 | 180 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010252 | Biological Process | auxin homeostasis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0048480 | Biological Process | stigma development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 243 aa Download sequence Send to blast |
MAGFFSLGGG GGGAGGGTQS GRATTSNNQE EESQNPNIGI SPENWFLYRN NQDEDNIPYN 60 KGFELWQYHQ QQQQQRHNHP AAVVQDLYPL DIGGGSSSSH HHGRLNLTSD EPNPRSGAAG 120 GFVMMRSSGS GGGGSGISCQ DCGNQAKKDC AHMRCRTCCK SRGYNCPTHV KSTWVPAAKR 180 RERQQQLASL QQQQQHQQNQ QQQQQHQLQL HHHHHRENTK RHRENPSSGN FRLKPSKEFF 240 RNF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator involved in the transcriptional regulation of terpene biosynthesis in glandular trichomes (PubMed:24884371, PubMed:24142382). Binds to the promoter of the linalool synthase TPS5 and promotes TPS5 gene transactivation (PubMed:24884371, PubMed:24142382). Acts synergistically with MYC1 in the transactivation of TPS5 (PubMed:24884371). {ECO:0000269|PubMed:24142382, ECO:0000269|PubMed:24884371}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00613 | PBM | Transfer from AT3G51060 | Download |
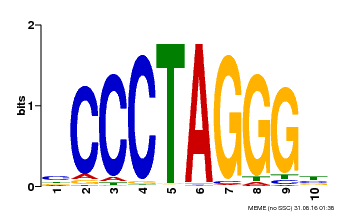
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016560706.1 | 1e-56 | PREDICTED: protein SHI RELATED SEQUENCE 1-like | ||||
| Swissprot | K4B6C9 | 2e-41 | EOT1_SOLLC; Protein EXPRESSION OF TERPENOIDS 1 | ||||
| TrEMBL | A0A0V0IDY8 | 4e-57 | A0A0V0IDY8_SOLCH; Uncharacterized protein | ||||
| STRING | XP_009763993.1 | 4e-55 | (Nicotiana sylvestris) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



