 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_12_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 362aa MW: 40167 Da PI: 9.3053 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 41.4 | 3.5e-13 | 63 | 111 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+++++g
cra_locus_12_iso_1_len_1626_ver_3 63 SRFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDIAAQRFRG 111
689***9788.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 97.7 | 7.6e-31 | 197 | 293 | 1 | 93 |
EEEE-..-HHHHTT-EE--HHH.HTT....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkan 72
f+k tpsdv+k++rlv+pk++ae+h +g++ +++ l +ed sg++W+++++y+++s++ vltkGW++Fvk++
cra_locus_12_iso_1_len_1626_ver_3 197 FEKSVTPSDVGKLNRLVIPKQHAEKHfplqSGNTSKGVLLNFEDMSGKVWRFRYSYWNSSKAXVLTKGWSRFVKEK 272
89****************************88999***************************************** PP
T--TT-EEEEEE-SSSEE..E CS
B3 73 gLkegDfvvFkldgrsefelv 93
+Lk+gD+v F+++ +++l+
cra_locus_12_iso_1_len_1626_ver_3 273 NLKAGDIVSFQRSTGIDKQLY 293
***********7655444455 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.05E-16 | 63 | 119 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 3.3E-8 | 63 | 111 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.90E-25 | 63 | 118 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.2E-19 | 64 | 119 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.954 | 64 | 119 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.6E-29 | 64 | 125 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 2.9E-39 | 190 | 302 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.88E-29 | 195 | 294 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.61E-28 | 196 | 285 | No hit | No description |
| Pfam | PF02362 | 1.0E-27 | 197 | 296 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.1E-22 | 197 | 300 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.817 | 197 | 300 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MDAGSCIDET TTSDTISTAP TSLSTLPVAK SPEISLCRVG SGASVILDSE GGVEAESRKL 60 PSSRFKGVVP QPNGRWGAQI YEKHQRVWLG TFNEEDEAAR AYDIAAQRFR GRDAVTNFKP 120 LSEISEEDDI ESAFLKFSSK AEIVDMLRKH TYNDELEQSK RNFGLNGGNF NKGIKDGFSN 180 PNNSGLGKNT EAREQLFEKS VTPSDVGKLN RLVIPKQHAE KHFPLQSGNT SKGVLLNFED 240 MSGKVWRFRY SYWNSSKAXV LTKGWSRFVK EKNLKAGDIV SFQRSTGIDK QLYIDWKPRN 300 TAAAIAAPIQ PVQTVRLFGV NIVSNVVVVD NTNITCSTGK RVREIEFLAF DCSKKPRIID 360 AL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 1e-56 | 194 | 300 | 11 | 118 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
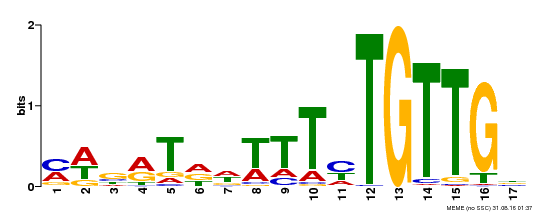
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018823697.1 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1-like | ||||
| Swissprot | Q9ZWM9 | 1e-144 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | W8CWZ4 | 0.0 | W8CWZ4_OLEEU; Tempranillo | ||||
| STRING | VIT_01s0011g03070.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2958 | 24 | 53 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



