 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_1271_iso_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 520aa MW: 57718.1 Da PI: 6.4632 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21 | 8.7e-07 | 270 | 291 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C++Cgk F+r nL+ H+r H
cra_locus_1271_iso_3_len_1945_ver_3 270 FCTICGKGFKRDANLRMHMRGH 291
6*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.47E-5 | 267 | 294 | No hit | No description |
| SMART | SM00355 | 0.0026 | 269 | 291 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.03 | 269 | 296 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.7E-6 | 270 | 320 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 271 | 291 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 58 | 318 | 351 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 31 | 356 | 378 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MDPDERQSAK AWTPSGNDVL KNLSMDNQSF IDFGLQQKWE DCSYLGYQSR IEQGFSKFSH 60 QSENSSSVDG SSNSQSKIQG RQDSRMQEAS NSNKMPDWDP KGMLSNLSFL EQKIHQLQGL 120 VQLIVGRRGQ VENQSNELLV QQQQLIIADL TSIIVQLIST AGSLLPSAKH TLSSVSPYVP 180 QFRQFGGSMV PSGTNASSGG GLPQNTVNKV EDQSQEIDLT GHHGTEQNFA VEEHEMKDED 240 DGEEGENLPP GSYEILQLEK EEILAPHTHF CTICGKGFKR DANLRMHMRG HGDEYKTPAA 300 LAKPHKESSS EPALIKRYSC PYVGCKRNKD HKKFQPLKTI LCVKNHYKRT HCDKSYTCSR 360 CHTKKFSVVA DLKTHEKHCG RDKWLCSCGT TFSRKDKLFG HIALFQGHTP AIPMDDSKGS 420 AGTSDGLQSS EAVNKIEQLD FSAQSNIPSG SCFENVMDGK GSVDDPVNYF SPLNFDPSNL 480 SGYQEFPRPS FEDSESSFSF LLSGSAQYDQ KSGRYSGPVI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
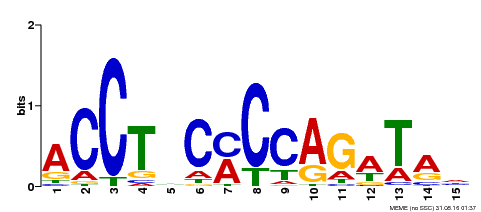
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027094877.1 | 0.0 | protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Refseq | XP_027094901.1 | 0.0 | protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X1 | ||||
| Swissprot | Q9C8N5 | 0.0 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A5BSF5 | 0.0 | A5BSF5_VITVI; Uncharacterized protein | ||||
| STRING | XP_008237278.1 | 0.0 | (Prunus mume) | ||||



