 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_12315_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 509aa MW: 55856 Da PI: 7.1547 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 49.8 | 6.1e-16 | 328 | 374 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ K +Ka++L +A+eY+ksLq
cra_locus_12315_iso_1_len_2437_ver_3 328 VHNLSERRRRDRINEKMRALQELLPHS-----NKSDKASMLDEAIEYMKSLQ 374
6*************************7.....59*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 1.09E-19 | 322 | 386 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.815 | 324 | 373 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.07E-17 | 327 | 378 | No hit | No description |
| Pfam | PF00010 | 2.1E-13 | 328 | 374 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 3.6E-19 | 328 | 383 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.3E-17 | 330 | 379 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 509 aa Download sequence Send to blast |
XAMNPCFPDW NIEAELPVLH QKKPFGLEHE LVELLWENGQ VVLHSQTHRK SGPAGVPDHP 60 FDSSTRQLNN NNNSNNKQAT TSCGNPTATL IQDNDTVSWI HCQVDDPFDK EFSSNFLPVI 120 PLPTPQLEDP PHKQQIQHCR THPSTNFGPN PLPAPRFQSD SNNVRGGVSK CPNVLLKADL 180 GSSNSGPRPS SHMIIGEVRE YSMRTVGSSH CGSNQVAMDA DTSRASSCGI GNNDLSAAAV 240 VKDYEGKLGS QNYERGERET HELANTSSSG GSGTSFGRTC TATDSLKRKS RDAEDSECQS 300 EAAVLESEAR KKPASKSGIA RKSRAAEVHN LSERRRRDRI NEKMRALQEL LPHSNKSDKA 360 SMLDEAIEYM KSLQLQLQMM WMRSGMAPLM FPGVQHYMSR LGMGIGPPTL PPVPNPMHLS 420 RLPLVDQTMT VASAPNQAAL CPTSVINPIN YQNQLQNSNF SEQYATYMGF HHPMQTSSQP 480 INVFGFNSNM TQQNHNFAPP SNTNGSSAG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 332 | 337 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
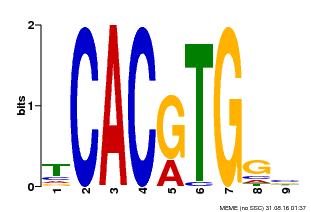
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027061311.1 | 1e-172 | transcription factor PHYTOCHROME INTERACTING FACTOR-LIKE 13-like isoform X1 | ||||
| Refseq | XP_027061312.1 | 1e-172 | transcription factor PHYTOCHROME INTERACTING FACTOR-LIKE 13-like isoform X1 | ||||
| Refseq | XP_027061313.1 | 1e-172 | transcription factor PHYTOCHROME INTERACTING FACTOR-LIKE 13-like isoform X1 | ||||
| TrEMBL | A0A0N7HUP6 | 0.0 | A0A0N7HUP6_CATRO; Phytochrome interacting factor 5-like protein | ||||
| STRING | XP_006485007.1 | 1e-167 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |



