 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_11227_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 295aa MW: 33444 Da PI: 6.6272 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.7 | 4.1e-18 | 75 | 129 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
rk+ +++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
cra_locus_11227_iso_5_len_1205_ver_3 75 RKKGRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 129
67778999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129 | 2e-41 | 75 | 167 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydal 73
+kk rl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr+++a+
cra_locus_11227_iso_5_len_1205_ver_3 75 RKKGRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDILKRQFEAI 147
6999********************************************************************* PP
HD-ZIP_I/II 74 keenerLekeveeLreelke 93
k+en++L++++++L++e+++
cra_locus_11227_iso_5_len_1205_ver_3 148 KAENDALQAQNQKLHAEIMA 167
***************99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.03E-19 | 64 | 133 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.491 | 71 | 131 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.6E-17 | 73 | 135 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.8E-15 | 75 | 129 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.78E-15 | 75 | 132 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-19 | 76 | 138 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.2E-5 | 102 | 111 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 106 | 129 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.2E-5 | 111 | 127 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.8E-17 | 131 | 171 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MLQTSHEDEH PHHQHPPTSL STPILSQDFH GVASFLGKRS MSFSGVDICD QDGVGGGGGH 60 GGEDDLSDDG SNWERKKGRL NMEQVKTLEK NFELGNKLEP ERKMQLARAL GLQPRQIAIW 120 FQNRRARWKT KQLEKDYDIL KRQFEAIKAE NDALQAQNQK LHAEIMALKN REPTESINLN 180 KETEGSCSNR SENSSDNIKL DISRTTPAID SHPQTSRPFF PSSLIRPNTA QQLFQTTSSS 240 SRPPPADHHH LIQCQNNNNN NPTATVKEES SLCNMFCTID DQTGFWPWLE QHHFN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 123 | 131 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
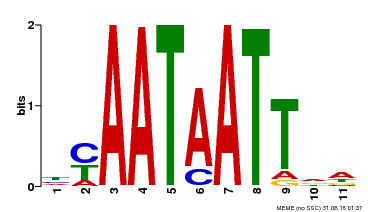
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027151684.1 | 1e-139 | homeobox-leucine zipper protein ATHB-13 | ||||
| Swissprot | Q8LC03 | 1e-111 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A1DR78 | 0.0 | A1DR78_CATRO; DNA-binding protein (Fragment) | ||||
| STRING | XP_002520111.1 | 1e-138 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA848 | 24 | 98 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



