 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_10685_iso_4 | ||||||||
| Common Name | myc1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 271aa MW: 29694.9 Da PI: 5.677 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 33.3 | 9e-11 | 154 | 201 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++YI+sLq
cra_locus_10685_iso_4_len_1403_ver_3 154 SHSLAERARREKISERMKILQDLVPGC----NKVIGKALVLDEIINYIQSLQ 201
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.26E-10 | 148 | 205 | No hit | No description |
| SuperFamily | SSF47459 | 9.55E-18 | 148 | 217 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 5.4E-18 | 150 | 218 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.828 | 150 | 200 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.9E-8 | 154 | 201 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-10 | 156 | 206 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048446 | Biological Process | petal morphogenesis | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 271 aa Download sequence Send to blast |
MDQQASYNLA GDTVPYNLAE IWPFPMNSAT AATSFHNNNN AANSLAAAAA AGISMNMNVN 60 LSVNRDHDPM VMDQAPNLNG GVRRRREDDD SAKGVSTSND ANAMNEGDNK RLKTGGSNEN 120 HESKAEGEET AKPAEPPKQD YIHVRARRGQ ATDSHSLAER ARREKISERM KILQDLVPGC 180 NKVIGKALVL DEIINYIQSL QRQVEFLSMK LEAVNSRLSP GIEGFPSKEF GQPPYDPSGM 240 AFGSQSSREY GRDTSPEWLH MQIGGGFERT T |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that forms a ternary complex with RSS3 and TIFY11A/JAZ9 to negatively regulate jasmonate-responsive genes. {ECO:0000269|PubMed:23715469}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00207 | DAP | Transfer from AT1G59640 | Download |
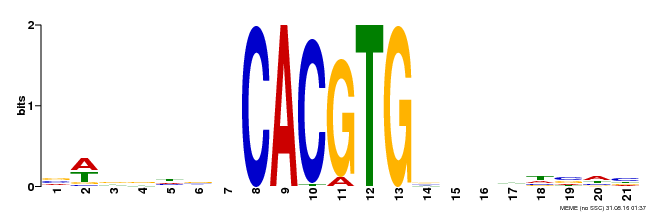
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027153368.1 | 1e-138 | transcription factor BHLH094 isoform X1 | ||||
| Swissprot | Q69WS3 | 3e-77 | BH094_ORYSJ; Transcription factor BHLH094 | ||||
| TrEMBL | Q71SQ1 | 0.0 | Q71SQ1_CATRO; MYC1 | ||||
| STRING | XP_009776012.1 | 1e-109 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1287 | 24 | 71 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



