 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_10685_iso_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 251aa MW: 27635.9 Da PI: 6.7997 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 33.4 | 7.9e-11 | 154 | 201 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++YI+sLq
cra_locus_10685_iso_3_len_1660_ver_3 154 SHSLAERARREKISERMKILQDLVPGC----NKVIGKALVLDEIINYIQSLQ 201
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 1.46E-9 | 148 | 205 | No hit | No description |
| SuperFamily | SSF47459 | 8.24E-18 | 148 | 217 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.7E-18 | 150 | 218 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.828 | 150 | 200 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.4E-8 | 154 | 201 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-10 | 156 | 206 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048446 | Biological Process | petal morphogenesis | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 251 aa Download sequence Send to blast |
MDQQASYNLA GDTVPYNLAE IWPFPMNSAT AATSFHNNNN AANSLAAAAA AGISMNMNVN 60 LSVNRDHDPM VMDQAPNLNG GVRRRREDDD SAKGVSTSND ANAMNEGDNK RLKTGGSNEN 120 HESKAEGEET AKPAEPPKQD YIHVRARRGQ ATDSHSLAER ARREKISERM KILQDLVPGC 180 NKVIGKALVL DEIINYIQSL QRQVEFLSMK LEAVNSRLSP GIEGFPSKEV SFTCVSLIAK 240 RRFTLELHVT W |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the control of petal size, by interfering with postmitotic cell expansion to limit final petal cell size. {ECO:0000269|PubMed:16902407}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00207 | DAP | Transfer from AT1G59640 | Download |
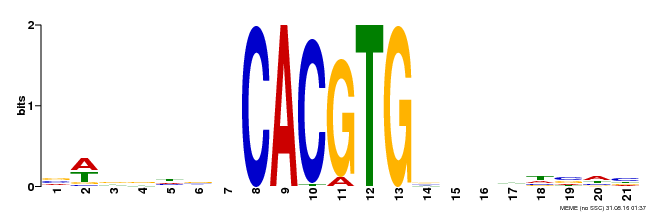
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Isoform 1 is up-regulated by PI/AP3, SEP2, SEP3 and AP1, but repressed by AG. {ECO:0000269|PubMed:16902407}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027153368.1 | 1e-110 | transcription factor BHLH094 isoform X1 | ||||
| Refseq | XP_027153369.1 | 1e-110 | transcription factor BHLH089 isoform X2 | ||||
| Swissprot | Q0JXE7 | 4e-71 | BPE_ARATH; Transcription factor BPE | ||||
| TrEMBL | Q71SQ1 | 1e-167 | Q71SQ1_CATRO; MYC1 | ||||
| STRING | XP_009776012.1 | 2e-87 | (Nicotiana sylvestris) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



