 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_10417_iso_4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 838aa MW: 96591.5 Da PI: 8.1519 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 86.1 | 5.5e-27 | 198 | 301 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrt 60
+fY+eYA+++GFs+ +++s++sk+++e+++++f Cs++g+++e +k+ e++ +ra +t
cra_locus_10417_iso_4_len_2511_ver_3 198 SFYQEYARSTGFSTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKSvnrprsrqgnrqdPENATGRRACAKT 270
5******************************************9999999999888776655666699***** PP
FAR1 61 gCkaklkvkkekdgkwevtkleleHnHelap 91
+Cka+++vk++ dgkw ++++e+eHnHel p
cra_locus_10417_iso_4_len_2511_ver_3 271 DCKASMHVKRRPDGKWIIHRFEKEHNHELLP 301
****************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.8E-25 | 198 | 301 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 6.3E-26 | 399 | 491 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.507 | 679 | 715 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 4.1E-4 | 686 | 712 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 8.3E-8 | 690 | 717 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 838 aa Download sequence Send to blast |
XLESGREGSV VVEDKGRRTR AVKRLIGCGW FWMSRNERVK ETEIFQAPNP NXQKKDNWET 60 SSPRPAPRVE FADSFAINNT TTLPARASES XQQLKGYLPG NLTLKKRYVM DIDLRLPSGE 120 QDKEDEEANG IAHMLDGDEE KPHNVDRLNG SMVDIGDKIN AENGGDIIPI TDMLDSKEDI 180 LLEPLAGMEF ESLGAAYSFY QEYARSTGFS TAIQNSRRSK TSREFIDAKF ACSRYGTKRE 240 YEKSVNRPRS RQGNRQDPEN ATGRRACAKT DCKASMHVKR RPDGKWIIHR FEKEHNHELL 300 PAQAVSEQTR RMYAAMAKQF AEYKNLVGLK SDSRSPFEKG RNPTIESGEA NLLLDFFTQM 360 QSINSNFYYA IEVGEDQRLK NFFWVDSKSR YDYINFSDVV SFDTTYIRNK YKMPLALFIG 420 VNQHYQFMLL GCALISDESS ATYSWVMRTW LRAMGSQAPK IIITDQDKVL KSVVSEIFSS 480 SLHFFFLWHI LGKVTETLSH VIKQNESFML KFEKCIYRSW TDEEFEKRWL KMVDKFELKE 540 NELIHTLYED RSRWVPTFVR DAFLAGMSTG QRSECVNSFF DKYLHKKTTI QEFVKQYDAI 600 LQDRYEEEAK ADSDTWNKQP ALKSPSPFEK HVANIYTHAV FKKFQIEVLG AVACIPKRER 660 EEETTTTFRV QDFEKNQEFA VTLNELKSEV SCICRLFEFK GFLCRHAMIV LQICGISSIP 720 SHYILKRWTK DAKGRYLMAE GSELVQSRVQ RYNDLCERAL RLAEEGSLST ESYNFALRAL 780 DDAFGNCVSA NNSNKNLAEA GPSATPGLLC MEEDEQSRSM NKTNKTNKKN NPTKKRKV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
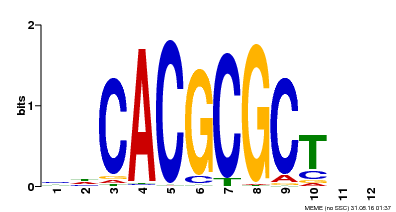
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027061868.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A1S4AZK2 | 0.0 | A0A1S4AZK2_TOBAC; protein FAR-RED ELONGATED HYPOCOTYL 3-like isoform X3 | ||||
| TrEMBL | A0A1U7YPD6 | 0.0 | A0A1U7YPD6_NICSY; protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X3 | ||||
| STRING | XP_009804946.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |



