 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cre16.g674890.t2.1 | ||||||||
| Common Name | CGLD5B, CHLREDRAFT_205920 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Chlamydomonadaceae; Chlamydomonas
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 656aa MW: 63885.4 Da PI: 6.6305 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.3 | 1.1e-19 | 375 | 425 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
++y+GVr+++ +g+++AeIrdp + r +lg+++taeeAa a+++a+++++g
Cre16.g674890.t2.1 375 CKYRGVRQRP-WGKYAAEIRDPH----KgCRLWLGTYDTAEEAALAYDKAAREIRG 425
79********.**********73....359************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.22E-31 | 375 | 435 | No hit | No description |
| Pfam | PF00847 | 1.9E-13 | 375 | 425 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.1E-31 | 376 | 434 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.54E-22 | 376 | 433 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 1.5E-38 | 376 | 439 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.42 | 376 | 433 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.7E-10 | 377 | 388 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.7E-10 | 399 | 415 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008219 | Biological Process | cell death | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0042991 | Biological Process | transcription factor import into nucleus | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048825 | Biological Process | cotyledon development | ||||
| GO:0051707 | Biological Process | response to other organism | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 656 aa Download sequence Send to blast |
MMHLDAKAKH SLLALDAASL GLALPGTSPP AAPLPVPGVK PPGAAGVAAL PPGSLPFGAA 60 TQFGQVPFFA GLPFPLPGGV PPPLPGLPPL SSAAAAAAAA AHAHAAAAAH AAAAMTIAQA 120 AAAAEAAPGK GRIGAIAASA AAATDPNIAV PTPPIAASPP TEPSDGSSPA TSHGSKDGAA 180 GDAVKTEPGA NTAAPGGALS SLGAIAAAAQ AATAGMAVPP SLWLGQPGSA GSAFQALAMP 240 MPPFPPNLAR TSVGGDGSVQ PGTSAPAHSD APSAPAPSGR PPLAPGMPGN LPSSLADHPA 300 AGICSTSYRS SGSHLGMPGG ALGASPPSLG TSPAAIKLPG RRRTSSTNLL SSSLGAKASS 360 SVPSLSVKPD KSGACKYRGV RQRPWGKYAA EIRDPHKGCR LWLGTYDTAE EAALAYDKAA 420 REIRGPRAVV NFPNVNHANL PAPQPHGDDE SGAWDPLGTT SLGTSPVSSG FIPGSSPLMG 480 GSAPVRHGMH PHHHLHGGAF SHHPGVAPSS GLGLRSTFGG YPVRREPTEP IAEGESDVDM 540 DDGEEGEDGD GDDMDEGVGP RSGRARMKVN VVTGVGGRTR PSRPAAAAAV AAAAAAAAGG 600 GVDLEDSDGA PPPPQPSMAG GASRGGGGKR NQVSMDVEDE LAELADALLL LHESA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 7e-23 | 377 | 433 | 6 | 63 | ATERF1 |
| 3gcc_A | 7e-23 | 377 | 433 | 6 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Cre.11444 | 0.0 | normal conditions| nutrient deficiency | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00109 | PBM | Transfer from AT3G16770 | Download |
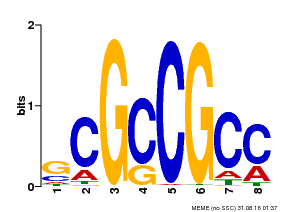
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cre16.g674890.t2.1 |
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001699213.1 | 0.0 | AP2-domain transcription factor | ||||
| TrEMBL | A0A2K3CU11 | 0.0 | A0A2K3CU11_CHLRE; Uncharacterized protein | ||||
| STRING | EDO98853 | 0.0 | (Chlamydomonas reinhardtii) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16770.1 | 2e-18 | ethylene-responsive element binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cre16.g674890.t2.1 |
| Entrez Gene | 5724763 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



