 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cre08.g358534.t1.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Chlamydomonadaceae; Chlamydomonas
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 839aa MW: 83504.6 Da PI: 4.7359 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 54.1 | 2.2e-17 | 229 | 258 | 1 | 30 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyy 30
C+ Cg+t+Tp+WR+gp g+ktLCnaCG++y
Cre08.g358534.t1.1 229 CVECGATQTPQWREGPAGPKTLCNACGVRY 258
****************************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51038 | 8.166 | 37 | 159 | IPR001025 | Bromo adjacent homology (BAH) domain |
| SMART | SM00401 | 1.3E-12 | 223 | 277 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 12.479 | 223 | 258 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 5.7E-12 | 226 | 263 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 3.9E-14 | 226 | 261 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 6.87E-12 | 228 | 258 | No hit | No description |
| Pfam | PF00320 | 8.3E-15 | 229 | 258 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 229 | 254 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MQKATIEFFW LGAPQNAASV HESADRVYYT GFNLNGVTFQ AGEYVLLHPE EEASLHYVGK 60 LVKCFHDKSG AYGDPHCVEV AWFERAVVLE AGADLKELVE LAETDINPIG CISGKAYVLK 120 ASSLEEASQL AQASVPAHCE WFWCARALRA GVLLPCSELA DPAGASACGQ QAALPPQLSG 180 KRSLTGGQGS AEGAADSEEA EAKRSRLQPE EMPTAPVRRT GKLVSGRTCV ECGATQTPQW 240 REGPAGPKTL CNACGVRYVR AQQRANKRAV SMGSSRPSGG YGGRQSRASK SARAEPACGP 300 RGGARAAASA AAVEEVPQRP LRQAALLAAN KTAQYARTGV FPQSATELHQ HQQQPYGGLC 360 TLGELELVGR PGTPVCPSAT ASAGASACAD SLPEGVTVEV EVVVDGGDMD GGCCGGGAVS 420 PAMGLVAPFA ALPGVPGGYP QLQQFQQHHH QALTHMGLLP GAGCSPERPA AGTSGLTHMG 480 GAAAVQGMVP LAGHVVLLGA AAAPAAAAAA ADSTPSTSGT SGSGLAAGAG GDCCASELPA 540 AAMPLQHHLL HPHVMSVPVS PSAHQPAATA ACDLFVCGDE DEAGAGSHMG LYGAGQSGLL 600 GAGEDEEAFG SLVGAEVDFG LGAGSGDGSA ALFDVLLQPG GPGGHCCEGD LDGDLGVGVC 660 DLPDVAARAA PYCTDLAPGT AARQLLDAAP LLLMAGDVPL GCGAGSSCGT VAVAGGAEAE 720 AAVRQLVGLG AEAGRAATEA LAASEAAAAV EQAAEAHAEA ARRARDAAAC NLERLREAIC 780 GIAGPEAAAA VALDCSGLIE LPHSLGGGDG GAVCAGGLLG MGLGSSLFDD ASLVAVHS* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Cre.12369 | 0.0 | normal conditions| nutrient deficiency | ||||
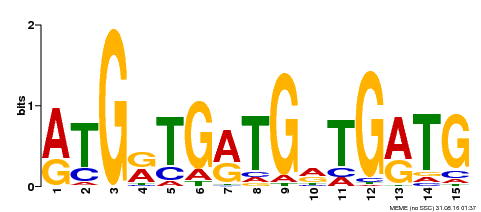
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00377 | DAP | Transfer from AT3G24050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cre08.g358534.t1.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001703730.1 | 0.0 | predicted protein | ||||
| TrEMBL | A0A2K3DG76 | 0.0 | A0A2K3DG76_CHLRE; Uncharacterized protein | ||||
| STRING | EDO96370 | 0.0 | (Chlamydomonas reinhardtii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 | Representative plant | OGRP68 | 17 | 287 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24050.1 | 2e-13 | GATA transcription factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cre08.g358534.t1.1 |



