 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_55.70 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 331aa MW: 36890.7 Da PI: 4.6982 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59 | 8e-19 | 58 | 111 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
evm.model.supercontig_55.70 58 KKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 111
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.8 | 1.1e-41 | 57 | 148 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLek 82
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk++y+alk + ++L++
evm.model.supercontig_55.70 57 EKKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKANYEALKLNYDALQR 138
69******************************************************************************** PP
HD-ZIP_I/II 83 eveeLreelk 92
++e+L +e++
evm.model.supercontig_55.70 139 DNEALMKEVR 148
*****99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.57E-19 | 43 | 115 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.265 | 53 | 113 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.4E-18 | 56 | 117 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 5.2E-16 | 58 | 111 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.24E-16 | 58 | 114 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-20 | 60 | 120 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.5E-6 | 84 | 93 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 88 | 111 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.5E-6 | 93 | 109 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.3E-18 | 113 | 154 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MKRSLGSSDS LGALMSICPN TDEHSPRNTH VYGREFQSML EGLDEEGCLE ESGHVGEKKR 60 RLSVDQVKAL EKNFEVENKL EPERKVKLAQ ELGLQPRQVA VWFQNRRARW KTKQLERDYG 120 VLKANYEALK LNYDALQRDN EALMKEVRGL KEKLNEEKTE SNLSVIKEEI ILPETDNNKT 180 LEPPPPSFSP PSPPASPLLP SAPLATTSDA TELNGVSAAS LFPTADFGSS DSDSSAILNE 240 ENNNNNNNNN NNSPNVAISL SAAGEFHHGL FISNSSSSPS CFPFSSRSTY NHTQFVKMEE 300 HNFFTGEEAC NFFSDEQAPT LQWYCAEQWN * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 105 | 113 | RRARWKTKQ |
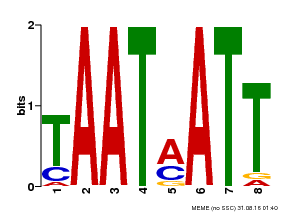
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_55.70 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021897528.1 | 0.0 | homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | A0A061DJ94 | 1e-139 | A0A061DJ94_THECC; Alanine--glyoxylate aminotransferase 2 isoform 1 | ||||
| STRING | evm.model.supercontig_55.70 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM545 | 28 | 143 | Representative plant | OGRP129 | 16 | 189 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 6e-67 | homeobox protein 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_55.70 |



