 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_183.29 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 297aa MW: 33977.9 Da PI: 6.8704 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56 | 6.8e-18 | 81 | 134 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
evm.model.supercontig_183.29 81 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 134
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 131.4 | 3.5e-42 | 80 | 172 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLe 81
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr+y+a+k++n++L+
evm.model.supercontig_183.29 80 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDLLKRQYEAIKADNDALQ 160
69******************************************************************************* PP
HD-ZIP_I/II 82 keveeLreelke 93
+++++L++e+++
evm.model.supercontig_183.29 161 AQNQKLHAEIMA 172
*******99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-19 | 57 | 137 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.92E-19 | 72 | 138 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 76 | 136 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 79 | 140 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.0E-15 | 81 | 134 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.57E-15 | 81 | 137 | No hit | No description |
| PRINTS | PR00031 | 1.2E-5 | 107 | 116 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 111 | 134 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.2E-5 | 116 | 132 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 9.0E-17 | 136 | 176 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 297 aa Download sequence Send to blast |
MSCNGMAFFP TNFMHQTPPP DHQDHHQPPL HTLLTSCTSQ DFHGVASFLG KRSMSFSGIE 60 VCEEGNGEDE LSDDGSQAGE KKRRLNMEQV KTLEKNFELG NKLEPERKMQ LARALGLQPR 120 QIAIWFQNRR ARWKTKQLEK DYDLLKRQYE AIKADNDALQ AQNQKLHAEI MALKSREPAE 180 SINLNKETEG SCSNRSENSS DIKLDISRTP AIDSPLPTHH HHHHHQPITS RALFPTSSSI 240 RPTTQLFQNS SSSSRPDNHL HQKMEQMVKE ESLTNMFCTI DDQSGFWPWL EQQHFN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 128 | 136 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
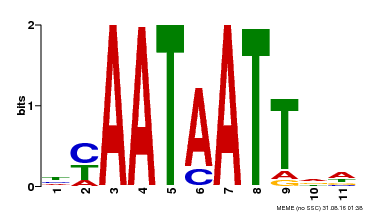
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_183.29 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021887869.1 | 0.0 | homeobox-leucine zipper protein ATHB-13-like | ||||
| Swissprot | Q8LC03 | 1e-120 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A1R3JEE3 | 1e-175 | A0A1R3JEE3_COCAP; Uncharacterized protein | ||||
| STRING | evm.model.supercontig_183.29 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2633 | 27 | 70 | Representative plant | OGRP129 | 16 | 189 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-115 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_183.29 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



