 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_14.18 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 280aa MW: 31462.3 Da PI: 6.5156 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.5 | 4.8e-18 | 72 | 124 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++ eq+++Le+ Fe +++ e++ +LAk lgL+ rq+ +WFqNrRa++k
evm.model.supercontig_14.18 72 KKRLNLEQVKALEKSFELGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWK 124
4568889*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.7 | 5.1e-41 | 70 | 160 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLek 82
ekk+rl+ eqvk+LE+sFe +kLeperK++la++Lglqprq+a+WFqnrRAR+ktkqlEkdye Lk++++alk++n++L++
evm.model.supercontig_14.18 70 EKKKRLNLEQVKALEKSFELGNKLEPERKMQLAKALGLQPRQIAIWFQNRRARWKTKQLEKDYEILKKQFEALKADNDALQA 151
69******************************************************************************** PP
HD-ZIP_I/II 83 eveeLreel 91
++++L++el
evm.model.supercontig_14.18 152 QNNKLNAEL 160
******998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 16.876 | 66 | 126 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.29E-19 | 66 | 128 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.5E-17 | 69 | 130 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.69E-16 | 71 | 127 | No hit | No description |
| Pfam | PF00046 | 2.7E-15 | 72 | 124 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-19 | 74 | 133 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.6E-6 | 97 | 106 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 101 | 124 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.6E-6 | 106 | 122 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.4E-14 | 126 | 166 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 280 aa Download sequence Send to blast |
MAFPSSHGFM FQPLDDSPSP TSLPSCPPHL FHGGGGFMMK RSMSFSGVEK REEIHGGDDE 60 LSDDGLQGGE KKKRLNLEQV KALEKSFELG NKLEPERKMQ LAKALGLQPR QIAIWFQNRR 120 ARWKTKQLEK DYEILKKQFE ALKADNDALQ AQNNKLNAEL LALKNKDCNE MNVKKEADQG 180 SWSNVSDSSC DINLDISRTA AVVSSSPVSS SQLSGRNFLS SSSSIRPPTS MTQLLQASSR 240 SDLHCQMIQQ EETFCNMFNP MEDQQQGFWP WSDQQQNFH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 118 | 126 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
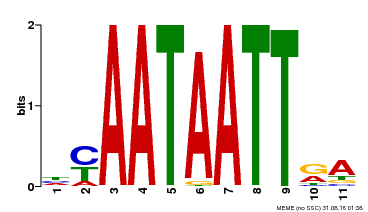
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_14.18 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021905273.1 | 0.0 | homeobox-leucine zipper protein ATHB-20-like | ||||
| Swissprot | Q8LAT0 | 8e-93 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A1R3I9H9 | 1e-132 | A0A1R3I9H9_COCAP; Uncharacterized protein | ||||
| STRING | evm.model.supercontig_14.18 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3095 | 26 | 66 | Representative plant | OGRP129 | 16 | 189 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 4e-73 | homeobox protein 20 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_14.18 |



