 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm.model.supercontig_136.16 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Caricaceae; Carica
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 389aa MW: 42454.9 Da PI: 8.853 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 94.7 | 6.7e-30 | 107 | 164 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK v+g+ef rsYY+Ct+++C vkk++ers+ d+++++++Y g+H+h+k
evm.model.supercontig_136.16 107 EDGYNWRKYGQKLVRGNEFIRSYYKCTHPKCLVKKQLERSH-DGRITDTVYFGQHDHPK 164
7****************************************.***************85 PP
| |||||||
| 2 | WRKY | 102.4 | 2.6e-32 | 279 | 337 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+++prsYYrC+s+gC+vkk+ver+++dpk++++tYeg+H+h+
evm.model.supercontig_136.16 279 VNDGYRWRKYGQKLVKGNPHPRSYYRCSSPGCRVKKHVERTSHDPKLLTTTYEGHHDHD 337
58********************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 1.7E-25 | 103 | 165 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.07E-23 | 104 | 165 | IPR003657 | WRKY domain |
| SMART | SM00774 | 3.0E-33 | 106 | 164 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.0E-22 | 107 | 163 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 22.641 | 107 | 165 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 1.0E-35 | 265 | 339 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 4.84E-29 | 271 | 339 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 34.612 | 274 | 339 | IPR003657 | WRKY domain |
| SMART | SM00774 | 3.2E-37 | 279 | 338 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 7.0E-26 | 280 | 336 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 389 aa Download sequence Send to blast |
MASSGQAATD EATSDKLPDG VSSVGGTREL QPISGSISVS ENHQEGNIPS ITPEKASVEP 60 DQGAVHSEGH VPPSKSKKEP KTPDRTAHAV QSDTEGISPV IREKGSEDGY NWRKYGQKLV 120 RGNEFIRSYY KCTHPKCLVK KQLERSHDGR ITDTVYFGQH DHPKPNSTPV AVGFVVSIIE 180 ERQNIASSVA KEKSSDAHGD TPHQTGPVGN LQVPAVASKD DVKDVLLGSN RVGDEPDTSD 240 DPGLKRRKKD NYNMEIISVE KQSSEPRIVV QTRSEVDIVN DGYRWRKYGQ KLVKGNPHPR 300 SYYRCSSPGC RVKKHVERTS HDPKLLTTTY EGHHDHDKPP ARTVTHNTAG SDVHKIPSNK 360 ESGSSVESGN KSKDSGTGCR ITPPSRKI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 1e-36 | 108 | 341 | 2 | 76 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
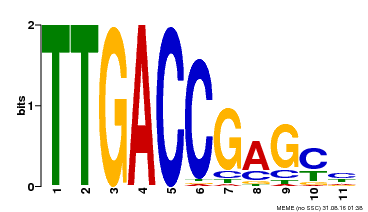
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | evm.model.supercontig_136.16 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021889975.1 | 0.0 | WRKY transcription factor 1 | ||||
| Swissprot | Q9SI37 | 3e-96 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | A0A061DMX7 | 1e-127 | A0A061DMX7_THECC; Zinc-dependent activator protein-1, putative isoform 1 | ||||
| STRING | evm.model.supercontig_136.16 | 0.0 | (Carica papaya) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7949 | 26 | 39 | Representative plant | OGRP14 | 17 | 875 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 1e-100 | zinc-dependent activator protein-1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm.model.supercontig_136.16 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



