| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 88.1 | 4.8e-28 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien rqvtfskRr+g+lKKA+EL +LCdaevaviifs+tgkl+e+ss
maker-scaffold03527-augustus-gene-0.20-mRNA-1 9 KRIENANSRQVTFSKRRAGLLKKAQELAILCDAEVAVIIFSNTGKLFEFSS 59
79***********************************************96 PP
|
| 2 | K-box | 57.8 | 4.7e-20 | 78 | 157 | 6 | 85 |
K-box 6 gksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskK 69
++++e + +++ ++el+ Lk+e+ Lq +q +llG+dL+ lkeLq++eqqL ++l +++kK
maker-scaffold03527-augustus-gene-0.20-mRNA-1 78 TSAVEYKAEKQESKELDVLKDEVARLQMKQLRLLGKDLTGVGLKELQHIEQQLSEGLLAVKEKK 141
4556777788999*************************************************** PP
K-box 70 nellleqieelqkkek 85
++ll++q+e+++ k +
maker-scaffold03527-augustus-gene-0.20-mRNA-1 142 EQLLMDQLEQSRLKCE 157
***********99865 PP
|
| Protein Features
? help Back to Top |
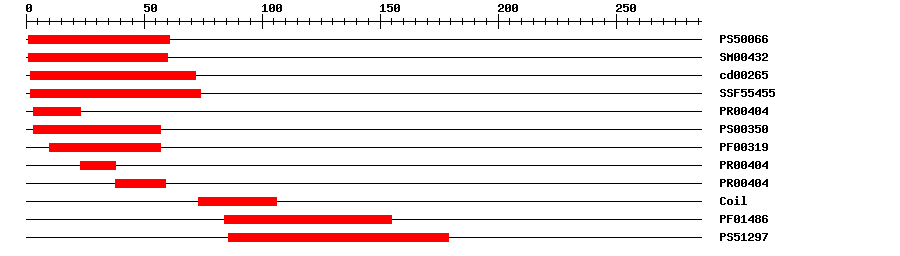 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
| PROSITE profile | PS50066 | 32.352 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 2.5E-41 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.76E-42 | 2 | 72 | No hit | No description |
| SuperFamily | SSF55455 | 1.44E-31 | 2 | 74 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 4.6E-29 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 8.4E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 4.6E-29 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 4.6E-29 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 8.7E-14 | 84 | 155 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 9.941 | 86 | 179 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0009553 | Biological Process | embryo sac development |
| GO:0010047 | Biological Process | fruit dehiscence |
| GO:0010094 | Biological Process | specification of carpel identity |
| GO:0010262 | Biological Process | somatic embryogenesis |
| GO:0010582 | Biological Process | floral meristem determinacy |
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated |
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated |
| GO:0048455 | Biological Process | stamen formation |
| GO:0048459 | Biological Process | floral whorl structural organization |
| GO:0048509 | Biological Process | regulation of meristem development |
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering |
| GO:0048833 | Biological Process | specification of floral organ number |
| GO:0060862 | Biological Process | negative regulation of floral organ abscission |
| GO:0060867 | Biological Process | fruit abscission |
| GO:0071365 | Biological Process | cellular response to auxin stimulus |
| GO:0080060 | Biological Process | integument development |
| GO:0080112 | Biological Process | seed growth |
| GO:2000692 | Biological Process | negative regulation of seed maturation |
| GO:0005634 | Cellular Component | nucleus |
| GO:0005737 | Cellular Component | cytoplasm |
| GO:0003677 | Molecular Function | DNA binding |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0042803 | Molecular Function | protein homodimerization activity |