 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | maker-scaffold00029-augustus-gene-1.37-mRNA-2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Fagaceae; Castanea
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 530aa MW: 60360.7 Da PI: 6.6243 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.2 | 2.5e-27 | 89 | 191 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............teker 52
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +e++
maker-scaffold00029-augustus-gene-1.37-mRNA-2 89 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSynrprarqnkpdSENAT 152
5******************************************999999999999888777777 PP
FAR1 53 rtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+r+ ++t+Cka+++vk++ dgkw+++++++eHnHel p
maker-scaffold00029-augustus-gene-1.37-mRNA-2 153 GRRSCSKTDCKASMHVKRRPDGKWVIHSFVKEHNHELLP 191
7999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.9E-25 | 89 | 191 | IPR004330 | FAR1 DNA binding domain |
| PROSITE profile | PS50966 | 8.988 | 231 | 267 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.3E-4 | 240 | 264 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.2E-7 | 242 | 269 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 530 aa Download sequence Send to blast |
MDIDLRLPSG EHDKEDEEPN AIDNILNGDE KLHNGDIEGE NMVDVGVEVR VEDGGDLNSP 60 TVDIGVFKED TNLEPLSGME FESHGEAYSF YQEYARSMGF NTAIQNSRRS KTSREFIDAK 120 FACSRYGTKR EYDKSYNRPR ARQNKPDSEN ATGRRSCSKT DCKASMHVKR RPDGKWVIHS 180 FVKEHNHELL PAQAKFQAEL LGVVACHPKR ERQDEAITTF RVQEFDRSQD FMVIWNEMKS 240 EISCICRLYE YKGYLCRHAM IVLQICGLSA IPSQYILKRW TKDAKSRHLL GEVSELIQSR 300 MQRYNDICQR AMKLSEEGSL SQESYSLAIR TLDEAFGTCV SMNNSSKSPT EVGTSDTHGL 360 LCIEEDNQSR SMGKTNKKKN PTKKRKMNSE SDVMTVSTQD SLQQMDKLSS RAVTLDSYYG 420 TPQSMQGMVQ LNLMAPTRDT YYGNQQTIQG LGQLNSIAPS HDGYYNAQQS MHGLGQMDFF 480 RTPTGFTYGI RDDPNAKRKQ CVEANVSVKN ELIMLCGWKD YGREPLLWHL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
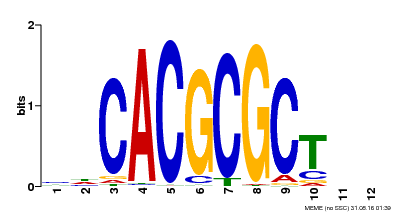
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023870500.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_023870501.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 1e-127 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A0A0KV81 | 0.0 | A0A0A0KV81_CUCSA; Uncharacterized protein | ||||
| STRING | VIT_03s0017g01780.t01 | 1e-162 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 1e-128 | far-red elongated hypocotyls 3 | ||||



