 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | augustus_masked-scaffold05815-abinit-gene-0.3-mRNA-1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Fagaceae; Castanea
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 294aa MW: 31891.6 Da PI: 9.555 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 40.4 | 6.8e-13 | 103 | 147 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ ++ + ++lG+g+W+ I+r ++Rt+ q+ s+ qky
augustus_masked-scaffold05815-abinit-gene-0.3-mRNA-1 103 PWTEEEHRTFLIGLEKLGKGDWRGISRNYVTTRTPTQVASHAQKY 147
7*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50158 | 8.532 | 3 | 18 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 18.218 | 96 | 152 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.05E-17 | 97 | 153 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.4E-18 | 99 | 151 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.6E-8 | 100 | 150 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-10 | 101 | 146 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.20E-8 | 103 | 148 | No hit | No description |
| Pfam | PF00249 | 1.8E-10 | 103 | 147 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MGRKCSHCGN IGHNSRTCTY RETFAGGLRL FGVQLDLSSS SFALKKSFSV DCLYQPSSSS 60 PTSPSSSLSS SRISIDENSD KTTIGYLSDG LICRPQERKK GIPWTEEEHR TFLIGLEKLG 120 KGDWRGISRN YVTTRTPTQV ASHAQKYFLR QASLNKKKRR SSLFDLVGSS SNILGHPASS 180 TNSKPSDAVP AEYCKTSSSQ HDTTLPLLGH SSSQQDAKSD NQETEDPQLP SSSKANVPVW 240 LYGLIDSQLK TSHAAKPSSS NASPNLELTL AAPRPLEQNK SSPAPLLFGP ITVT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 155 | 159 | KKKRR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
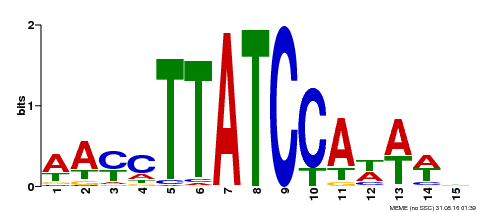
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023906593.1 | 0.0 | transcription factor MYBS3-like isoform X2 | ||||
| TrEMBL | A0A2N9EP74 | 1e-161 | A0A2N9EP74_FAGSY; Uncharacterized protein | ||||
| STRING | VIT_11s0016g02410.t01 | 1e-116 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3213 | 32 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56840.1 | 1e-53 | MYB_related family protein | ||||



