 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C024001P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 252aa MW: 28598.7 Da PI: 6.2624 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 88.4 | 3.9e-28 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien rqvtfskRr+g+lKKA+EL +LCdaevaviifs+tgkl+e+ss
MELO3C024001P1 9 KRIENANSRQVTFSKRRAGLLKKAQELAILCDAEVAVIIFSNTGKLFEFSS 59
79***********************************************96 PP
| |||||||
| 2 | K-box | 68.5 | 2.1e-23 | 86 | 172 | 14 | 100 |
K-box 14 aeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
+++ + e++ L++ei +Lq +q +llG+dL+ L +keLq+LeqqL+++l +++kK++ll+eq+e+++ +e+++ en++Lr++++e
MELO3C024001P1 86 EKREHDEVDILREEITTLQMKQLQLLGKDLTGLGFKELQNLEQQLNEGLLLVKEKKEQLLMEQLEQSRVQEQRAMLENETLRRQVNE 172
567889*****************************************************************************9976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 4.8E-42 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 32.464 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 3.27E-32 | 2 | 76 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 3.21E-43 | 2 | 76 | No hit | No description |
| PRINTS | PR00404 | 3.4E-29 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 6.9E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.4E-29 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.4E-29 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.4E-16 | 86 | 170 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 13.422 | 86 | 176 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 252 aa Download sequence Send to blast |
MGRGKIEIKR IENANSRQVT FSKRRAGLLK KAQELAILCD AEVAVIIFSN TGKLFEFSSS 60 GMKHTLARYN KCVESSDATV DVHKVEKREH DEVDILREEI TTLQMKQLQL LGKDLTGLGF 120 KELQNLEQQL NEGLLLVKEK KEQLLMEQLE QSRVQEQRAM LENETLRRQV NELRCLFPPV 180 DCPLPAYLEY CSLEQKNIGI RSPDMACNSE IERGDSDTTL HLGLPSHVYC KRKESERDTH 240 SNDSGAQIGI L* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6bz1_A | 3e-22 | 1 | 90 | 1 | 93 | MEF2 CHIMERA |
| 6bz1_B | 3e-22 | 1 | 90 | 1 | 93 | MEF2 CHIMERA |
| 6bz1_C | 3e-22 | 1 | 90 | 1 | 93 | MEF2 CHIMERA |
| 6bz1_D | 3e-22 | 1 | 90 | 1 | 93 | MEF2 CHIMERA |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the negative regulation of flowering, probably through the photoperiodic pathway. Acts as both an activator and a repressor of transcription. Binds DNA in a sequence-specific manner in large CArG motif 5'-CC (A/T)8 GG-3'. Participates probably in the regulation of programs active during the early stages of embryo development. Prevents premature perianth senescence and abscission, fruits development and seed desiccation. Stimulates the expression of at least DTA4, LEC2, FUS3, ABI3, AT4G38680/CSP2 and GRP2B/CSP4. Can enhance somatic embryo development in vitro. {ECO:0000269|PubMed:10318690, ECO:0000269|PubMed:10662856, ECO:0000269|PubMed:12226488, ECO:0000269|PubMed:12743119, ECO:0000269|PubMed:14615187, ECO:0000269|PubMed:15084721, ECO:0000269|PubMed:15686521, ECO:0000269|PubMed:17521410, ECO:0000269|PubMed:18305206, ECO:0000269|PubMed:19269998, ECO:0000269|PubMed:19767455, ECO:0000269|PubMed:8953767}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
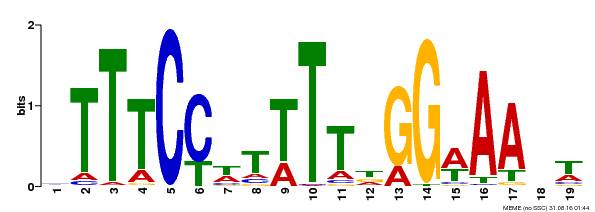
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin (2,4-D). Feedback loop leading to direct down-regulation by itself. {ECO:0000269|PubMed:15686521}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681835 | 2e-96 | LN681835.1 Cucumis melo genomic scaffold, anchoredscaffold00064. | |||
| GenBank | LN713258 | 2e-96 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008461826.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL15 isoform X1 | ||||
| Swissprot | Q38847 | 1e-84 | AGL15_ARATH; Agamous-like MADS-box protein AGL15 | ||||
| TrEMBL | A0A1S3CFG4 | 0.0 | A0A1S3CFG4_CUCME; agamous-like MADS-box protein AGL15 isoform X1 | ||||
| STRING | XP_008461826.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3582 | 32 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13790.1 | 2e-69 | AGAMOUS-like 15 | ||||



