 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C023853P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 324aa MW: 34444.4 Da PI: 9.0158 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 123.8 | 5.5e-39 | 75 | 135 | 2 | 62 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkkss 62
+e+ lkcprC+stntkfCy+nny+lsqPr+fCk+CrryWt+GGalrnvPvGgg+r+nk+++
MELO3C023853P1 75 PEAGLKCPRCESTNTKFCYFNNYNLSQPRHFCKTCRRYWTRGGALRNVPVGGGCRRNKRTK 135
67889*****************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 8.0E-34 | 59 | 128 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 1.9E-32 | 78 | 133 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 29.767 | 79 | 133 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 81 | 117 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009640 | Biological Process | photomorphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
MVFPPLPSYL DPPNWQPLTN HQTGGNDTAE DSGQVLLPPP PGGGGGGESA GGGSTGSIRP 60 ISMTDRARLA KIPQPEAGLK CPRCESTNTK FCYFNNYNLS QPRHFCKTCR RYWTRGGALR 120 NVPVGGGCRR NKRTKSRNRS KSPAASERQL LVGSTNSTAA VSLSTQPHLP FLSSLQNFST 180 YGLNLGIIPP QAPPTSGGSA AGGVDVHEFQ GDHWRLQQPQ QFPLLANDQQ PNLLYTFEPP 240 EGAMTRYNLG GFSRLGIEND IGMSMEGEAV KVEESKGMNL GKNFQFWVGG GGGNDVNAWS 300 GGGSGGSELH GFSSSSASHL LRQ* |
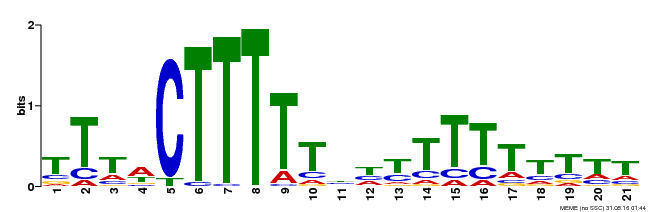
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00407 | DAP | Transfer from AT3G55370 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681897 | 0.0 | LN681897.1 Cucumis melo genomic scaffold, anchoredscaffold00063. | |||
| GenBank | LN713264 | 0.0 | LN713264.1 Cucumis melo genomic chromosome, chr_10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008461665.1 | 0.0 | PREDICTED: dof zinc finger protein DOF2.4 | ||||
| TrEMBL | A0A1S3CF93 | 0.0 | A0A1S3CF93_CUCME; dof zinc finger protein DOF2.4 | ||||
| STRING | XP_008461665.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF656 | 34 | 143 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G37590.1 | 6e-44 | DNA binding with one finger 2.4 | ||||



