 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C022532P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 873aa MW: 93910.8 Da PI: 5.9167 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 64.9 | 1.1e-20 | 166 | 221 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+++l L++rqVk+WFqNrR+++k
MELO3C022532P1 166 KKRYHRHTPQQIQELEAVFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 221
688999***********************************************999 PP
| |||||||
| 2 | START | 210.1 | 8.5e-66 | 374 | 600 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetl 82
ela++a++elvk+a+ +ep+W s e++n++e++++f++ + + +ea+r+sg+v+ ++ lve+l+d++ +W e+++ + +t+
MELO3C022532P1 374 ELALAAMDELVKMAQTDEPLWIGSLeggrEILNQEEYIRTFTPCIGmkpngFVTEASRESGMVIINSLALVETLMDSN-RWAEMFPcmiaRTTTT 467
5899**************************************9988999*9***************************.**************** PP
EEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--...-TTSEE-EESSEEEEEEEECTCEEEEEEEE CS
START 83 evissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe..sssvvRaellpSgiliepksnghskvtwve 168
+vis+g galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvSvd ++ p+ ss+ +++lpSg+++++++ng+skvtwve
MELO3C022532P1 468 DVISNGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSVDAMRETPTggGSSFGNCRRLPSGCVVQDMPNGYSKVTWVE 562
*************************************************************999******************************* PP
-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 169 hvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
h++++++++h+l+r+l++sg+ +ga++wv tlqrqce+
MELO3C022532P1 563 HAEYDDSQVHQLYRPLLSSGMGFGAQRWVTTLQRQCEC 600
************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 9.41E-21 | 148 | 223 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-22 | 153 | 223 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.2 | 163 | 223 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.3E-18 | 164 | 227 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 9.53E-19 | 165 | 223 | No hit | No description |
| Pfam | PF00046 | 3.0E-18 | 166 | 221 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 198 | 221 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.789 | 365 | 603 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.56E-33 | 367 | 600 | No hit | No description |
| CDD | cd08875 | 5.08E-125 | 369 | 599 | No hit | No description |
| Pfam | PF01852 | 1.3E-57 | 374 | 600 | IPR002913 | START domain |
| SMART | SM00234 | 2.7E-51 | 374 | 600 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.43E-23 | 628 | 798 | No hit | No description |
| SuperFamily | SSF55961 | 2.43E-23 | 825 | 865 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 873 aa Download sequence Send to blast |
MSFGGFLDGG GGGGGGGARI LADLPYTNNS TTNANNNPTG GIGGGGNMSS SAIAPPRLIT 60 QSLTKSMFNS PGLSLALVLI FFPLFFLRYS FRLFLCVVVP ISNGFENPQT NMDGGQGDLA 120 ARLPEGFEHN VGRRGREEEH ESRSGSDNMD GGSGDDQDAA DNPPRKKRYH RHTPQQIQEL 180 EAVFKECPHP DEKQRLELSR RLCLETRQVK FWFQNRRTQM KTQLERHENT LLRQENDKLR 240 AENMSIRDAM RNPICSNCGG PAIIGEISLE EQQLRIENAR LKDELDRVCA LAGKFLGRPI 300 SSLANSIAPP LPSSSLELGV GSNGFGSLTM ATSMPIGPDF GGGLSGNLAV VQAPARPTPG 360 MGLDRSVERS MLLELALAAM DELVKMAQTD EPLWIGSLEG GREILNQEEY IRTFTPCIGM 420 KPNGFVTEAS RESGMVIINS LALVETLMDS NRWAEMFPCM IARTTTTDVI SNGMGGTRNG 480 ALQLMHAELQ VLSPLVPVRE VNFLRFCKQH AEGVWAVVDV SVDAMRETPT GGGSSFGNCR 540 RLPSGCVVQD MPNGYSKVTW VEHAEYDDSQ VHQLYRPLLS SGMGFGAQRW VTTLQRQCEC 600 LAILMSSAVP IRDHTAITAG GRRSMLKLAQ RMTANFCAGV CASTVHKWNK LNAGSVDEDV 660 RVMTRKSVDD PGEPPGIVLS AATSVWLPVS PQRLFDFLRD ERLRSEWDIL SNGGPMQEMA 720 HIAKGQDHGN CVSLLRASAM NANQSSMLIL QETCIDAAGS LVVYAPVDIP AMHVVMNGGD 780 SAYVALLPSG FAIVPDGAVT GGLTATNGSS PSGGEGPQSQ RAAGGGSLLT VAFQILVNSL 840 PTAKLTVESV ETVNNLISCT VQKIKAALQC ET* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
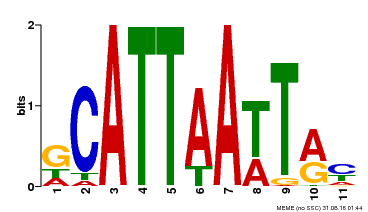
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681925 | 0.0 | LN681925.1 Cucumis melo genomic scaffold, anchoredscaffold00052. | |||
| GenBank | LN713265 | 0.0 | LN713265.1 Cucumis melo genomic chromosome, chr_11. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008460172.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A1S3CBG8 | 0.0 | A0A1S3CBG8_CUCME; homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| STRING | XP_008460172.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



