 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C019925P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 322aa MW: 35886.5 Da PI: 9.1585 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 49.8 | 7.5e-16 | 251 | 300 | 5 | 55 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkke 55
+r++r++kNRe+A rsR+RK+a++ eLe+kv Le+eN++L+k+ +el+k+
MELO3C019925P1 251 RRQKRMIKNRESAARSRARKQAYTNELENKVSRLEEENERLRKR-KELEKM 300
79****************************************55.777776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 7.0E-14 | 241 | 299 | No hit | No description |
| SMART | SM00338 | 1.9E-12 | 247 | 319 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.691 | 249 | 294 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 1.15E-22 | 251 | 305 | No hit | No description |
| Pfam | PF00170 | 3.0E-13 | 251 | 298 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 8.75E-11 | 251 | 296 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 254 | 269 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 322 aa Download sequence Send to blast |
MGIQTMGSQA GGDPNGKQSQ FQPLVRQNSL YSLTLDEVQN QLGDLGKPLI SMNLDELLKN 60 VWTAEANQTV GKDNEDNNIL ANQTSLQRQA SLSLNGALSK KTVDEVWRDI QQSKDSEEKK 120 SRERQPTLGE MTLEDFLVKA GVVAETSSNK KGAGPVVEID ANITPQFQQT QWMQYTQPQY 180 QSQQAAMMGV YMSGQPIPQP LHVGGGAVMD VPYVDNQLAL PTPLMGALSD TQASGRKRGA 240 PEDMIEKTVE RRQKRMIKNR ESAARSRARK QAYTNELENK VSRLEEENER LRKRKELEKM 300 LPLAPSPEPK YQLRRTSSAP F* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the embryo specification element and the ABA-responsive element (ABRE) of the Dc3 gene promoter. Could participate in abscisic acid-regulated gene expression during seed development. | |||||
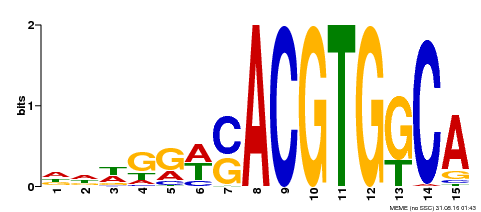
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00409 | DAP | Transfer from AT3G56850 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681821 | 0.0 | LN681821.1 Cucumis melo genomic scaffold, anchoredscaffold00040. | |||
| GenBank | LN713257 | 0.0 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008456642.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 2 isoform X1 | ||||
| Refseq | XP_008456643.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 2 isoform X1 | ||||
| Refseq | XP_008456644.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 2 isoform X1 | ||||
| Swissprot | Q9LES3 | 4e-96 | AI5L2_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 2 | ||||
| TrEMBL | A0A1S3C3Q8 | 0.0 | A0A1S3C3Q8_CUCME; ABSCISIC ACID-INSENSITIVE 5-like protein 2 isoform X1 | ||||
| STRING | XP_008456642.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1320 | 34 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G56850.1 | 1e-80 | ABA-responsive element binding protein 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



