 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C018642P2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1093aa MW: 121495 Da PI: 5.5526 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 182.4 | 5.2e-57 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqr 96
++ k+rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktvrE+hekLKvg+++vl+cyYah+een++fqr
MELO3C018642P2 20 IEAKHRWLRPAEICEILRNYTKFRIASEPPDRPSSGSLFLFDRKVLRYFRKDGHKWRKKKDGKTVREAHEKLKVGSIDVLHCYYAHGEENENFQR 114
4569******************************************************************************************* PP
CG-1 97 rcywlLeeelekivlvhylevk 118
r+yw+Lee+l +iv+vhylevk
MELO3C018642P2 115 RSYWMLEEHLMHIVFVHYLEVK 136
*******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.36 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.8E-80 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.2E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.2E-8 | 491 | 593 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 5.4E-6 | 507 | 580 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 9.02E-16 | 508 | 592 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.34E-15 | 687 | 802 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 15.804 | 691 | 773 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.24E-11 | 691 | 800 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-13 | 695 | 804 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF52540 | 2.02E-8 | 914 | 965 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.014 | 914 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.352 | 915 | 944 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 5.8E-4 | 916 | 935 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 937 | 959 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.615 | 938 | 962 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.8E-4 | 940 | 959 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1093 aa Download sequence Send to blast |
MADRGSYGLA PRLDIEQLLI EAKHRWLRPA EICEILRNYT KFRIASEPPD RPSSGSLFLF 60 DRKVLRYFRK DGHKWRKKKD GKTVREAHEK LKVGSIDVLH CYYAHGEENE NFQRRSYWML 120 EEHLMHIVFV HYLEVKGNRT NVGAVVETDE VSTSSQKSST KSSSYSSSHN QAASENADSP 180 SPTSTLTSFC EDADNEDTYQ ATSRFHSFPT SPKMGNGLLV NKPDAGQSNF YFPHSSSNNV 240 EGWSSVPAVD YVAQVQKDGL GGNSGDTSMM GSQKTLSSAS WEEILHQCTT GYQTVPSHVL 300 TSSIEPLSSG IVIGQENSTP DKILTSNSAI KEDFGNSLTM TSNWQVPFED NTLSFSKEHV 360 DHFPDLYSVC DIDSRLTAQK SHDATFGSGH EMFCAHPGKQ NEEILPNLEL QFKEGESYPA 420 MRLSSDNDMP KEGTISYSLT LKQSLIDGEE SLKKVDSFSR WVSKELGEVD DLHMHPSSGL 480 SWTTVECGDM VDDSSLSPSI SEDQLFSITA FSPKWTVTDL ETEVVVIGRF LGNNNGTNCH 540 WSCMFGEVEV PAEVLADGIL CCHAPPHSVG RVPFYVTCSN RVACSEVREF DYLAGSAQDV 600 DVTDIYTAGA TEELRMHLRF ERLLSLEPSD PSNDLSEGAL EKQNLIRELI TIKEEDDSYG 660 EDPNPQNDQI QHQSKEFLFV KLMKEKLYSW LIHKVIEGGK GPNILDGEGQ GVIHLAAALG 720 YDWAIRPIVA AGVSINFRDI NGWTALHWAA LCGRELTVVT LITLDASPGL MSDPSPEVPL 780 GIVPADLASI NGHKGISGFL AEAALTSYVS SISMAETVDD GVSDVSKTKA VQTVSERKAT 840 PVNDGFGDLS LKDSLTAVCN ATQAAGRIYQ ILRVQSFQRK KLSECGTDEF GSSDNSILSF 900 MKARARKSGL SNNPAHAAAV QIQKKFRGWR MRKEFLLIRQ RIVKIQAHVR GHQVRKQYRK 960 IVWSVGMIDK IILRWRRKGS GLRGFRSDAV AKDPPSLMAP PTKEDDYDFL KEGRRQTEER 1020 FQKALTRVKS MAQYPEGRDQ YRRLLTVVQK CRETKGSAMV VTSTSEEVIE GDDMIDIDTL 1080 LDDDALMSMT FD* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
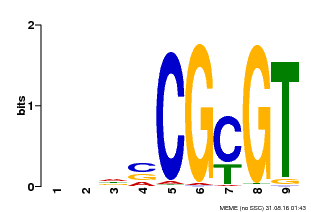
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681792 | 0.0 | LN681792.1 Cucumis melo genomic scaffold, anchoredscaffold00034. | |||
| GenBank | LN713255 | 0.0 | LN713255.1 Cucumis melo genomic chromosome, chr_1. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008455205.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1S3C137 | 0.0 | A0A1S3C137_CUCME; calmodulin-binding transcription activator 2 isoform X2 | ||||
| STRING | XP_008455204.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



