 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C016234P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 614aa MW: 72166.5 Da PI: 8.6436 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.7 | 1.7e-27 | 41 | 127 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk++ dg+w ++++ ++HnHel p
MELO3C016234P1 41 FYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESESG---NSRRPSVKKTDCKASMHVKRRPDGRWIIHEFIKDHNHELLP 127
***************************************9998877...7788899*******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.9E-25 | 41 | 127 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 5.4E-11 | 186 | 242 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.371 | 429 | 465 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 9.6E-6 | 438 | 464 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.3E-7 | 440 | 467 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 614 aa Download sequence Send to blast |
MQDTEGAIVS LPKKDILFEG DVDFEPHTGI EFESHEAAYT FYQEYAKSMG FTTSIKNSRR 60 SKKSKEFIDA KFACSRYGVT PESESGNSRR PSVKKTDCKA SMHVKRRPDG RWIIHEFIKD 120 HNHELLPALA YHFRIHRNVK LAEKNNIDIL HAVSRYLALD EGDAQILLEY FKRIQKENPY 180 FFYAIDLNEE QRLRNLFWVD AKTMGGKAPK VIITDQDKAL KLAIEEVFPN TRHCFALWHI 240 LEKIPETLAH VIKRHENFLA KFNKCIFKSW SDEQFDMRWW KMVTRFELQD DEWIQSLYDD 300 RRKWVPTYME DIFLAGMSTA QRSDSMNAFF DKYIHKKITL KEFLRQYGII LQNRYEEEVI 360 ADFDTLHKQP ALKSPSPWEK QMSTIYTHTI FKKFQVEVLG VVGCRMRKEI EDGTVTTFRV 420 QDCEKDEHFL VRWHKLNSEV SCFCRLFEYK GFLCRHALIV LQMLDFRSIP SQYILKRWTK 480 DAKSRQPVTE ETEFRQNRVQ RYNDLCKKAI ELSEEGSHSE ECYNIAIRTL VEALKNCVNI 540 NNSKSAPAES SVHAHGLREE EENQGSITTK ANKKKSTNRK RKVGFQLAFD KLLYLCSEQV 600 KHGSHVFLFA KDS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
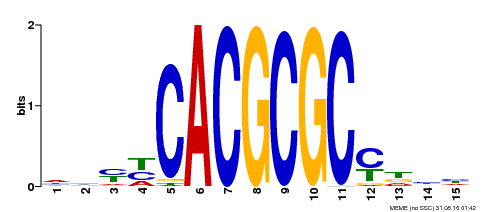
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681873 | 0.0 | LN681873.1 Cucumis melo genomic scaffold, anchoredscaffold00027. | |||
| GenBank | LN713261 | 0.0 | LN713261.1 Cucumis melo genomic chromosome, chr_7. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022136412.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X2 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A438BXP3 | 0.0 | A0A438BXP3_VITVI; Protein FAR-RED impared response 1 | ||||
| STRING | XP_009349572.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF21 | 14 | 477 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



