 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C010843P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 380aa MW: 41398.9 Da PI: 5.2411 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 50.9 | 2.7e-16 | 200 | 246 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn+ E+rRR+riN+++ L+ l+P++ K +Ka++L +A+eY+k+Lq
MELO3C010843P1 200 VHNMSEKRRRRRINEKMKALQNLIPNS-----NKTDKASMLDEAIEYLKQLQ 246
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 1.99E-18 | 192 | 250 | No hit | No description |
| SuperFamily | SSF47459 | 1.57E-20 | 193 | 256 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.2E-20 | 193 | 256 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.949 | 196 | 245 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 9.4E-14 | 200 | 246 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.5E-18 | 202 | 251 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010114 | Biological Process | response to red light | ||||
| GO:0010154 | Biological Process | fruit development | ||||
| GO:0010187 | Biological Process | negative regulation of seed germination | ||||
| GO:0048440 | Biological Process | carpel development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 380 aa Download sequence Send to blast |
MADLYGTSPS SDVVGLPHDP EDFSSFINHH LQTTTSSTSS PSCVSFKNKF MPLLHSQPPW 60 HSSPMFSRRP DSTSVPGMSC MLDFPEDRLH SSRVLNHALS DCSVRDVRFA DASYLAVNST 120 CGGVKLSDPG DFVKESSDNA FSSSGAVDSD TNAPLKRRGL SSENDLGDFS CDSEGGDLPE 180 VPSSTDLPRN SSKRSRSAEV HNMSEKRRRR RINEKMKALQ NLIPNSNKTD KASMLDEAIE 240 YLKQLQLQVQ MLSMRNGLSL QPMCLPGVLQ PIQLPQMGLD FDVGNAFLTS RRGIDTSSTR 300 NEGCPMQSTF NLTNKCNLSD QSIAIPSVPN TTTSETAFGF EPTIQGYNGE FNLSSDFKDG 360 EHAAQLDCTR TGKESTDAS* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 205 | 210 | KRRRRR |
| 2 | 205 | 211 | KRRRRRI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that plays a role in floral organogenesis. Promotes the growth of carpel margins and of pollen tract tissues derived from them. {ECO:0000269|PubMed:10225997, ECO:0000269|Ref.8}. | |||||
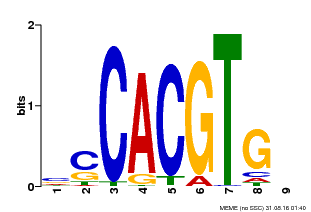
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00035 | PBM | Transfer from AT4G36930 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by the A class gene AP2 in the first whorl and by ARF3/ETT in gynoecium. {ECO:0000269|PubMed:11245574}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681823 | 0.0 | LN681823.1 Cucumis melo genomic scaffold, anchoredscaffold00014. | |||
| GenBank | LN713257 | 0.0 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008444604.1 | 0.0 | PREDICTED: transcription factor SPATULA-like | ||||
| Swissprot | Q9FUA4 | 2e-38 | SPT_ARATH; Transcription factor SPATULA | ||||
| TrEMBL | A0A1S3BA80 | 0.0 | A0A1S3BA80_CUCME; transcription factor SPATULA-like | ||||
| STRING | XP_008444604.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2276 | 33 | 79 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36930.1 | 1e-40 | bHLH family protein | ||||



