 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C009265P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 341aa MW: 36145.1 Da PI: 9.8407 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 123.8 | 5.8e-39 | 78 | 137 | 3 | 62 |
zf-Dof 3 ekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkkss 62
e+alkcprC+stntkfCy+nnysl+qPr+fCk+CrryWt+GGalrnvPvGgg+r+nk+++
MELO3C009265P1 78 EAALKCPRCESTNTKFCYFNNYSLTQPRHFCKTCRRYWTRGGALRNVPVGGGCRRNKRTK 137
7789*****************************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101447 | 3.92E-5 | 41 | 48 | No hit | No description |
| ProDom | PD007478 | 1.0E-36 | 61 | 135 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 1.5E-32 | 80 | 135 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 29.544 | 81 | 135 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 83 | 119 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MNFSSIPPYL DPSNWQQQVT HQVGTSSSTA VSSQLLPPPP PPPPPPPPPP PHGIGGAGSI 60 RPGSMAERAR MANIPMPEAA LKCPRCESTN TKFCYFNNYS LTQPRHFCKT CRRYWTRGGA 120 LRNVPVGGGC RRNKRTKGSS SKSPPVSSDR QQTSGSANSS SSAIASNNSG GLSPQIPPLG 180 RFMAPLHQQL SDFDIGSYSY GGGLSAPATA TGDLNFHLGN TNLGGGTSIG SLLGFDQQQW 240 RLQQQPPQFP FLSSLDPFEG GNGGGGEAPG TGWPMRPKVP SRNNLTQMGN SVKMEETPDQ 300 VNNNVGRQLI GNNEQYWSSG SMAWSDLSGF SSSSSTRNPL * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Probably involved in early processes for vascular development (PubMed:17583520). The PEAR proteins (e.g. DOF2.4, DOF5.1, DOF3.2, DOF1.1, DOF5.6 and DOF5.3) activate gene expression that promotes radial growth of protophloem sieve elements. Triggers the transcription of HD-ZIP III genes, especially in the central domain of vascular tissue (PubMed:30626969). {ECO:0000250|UniProtKB:Q9M2U1, ECO:0000269|PubMed:17583520, ECO:0000269|PubMed:30626969}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00295 | DAP | Transfer from AT2G37590 | Download |
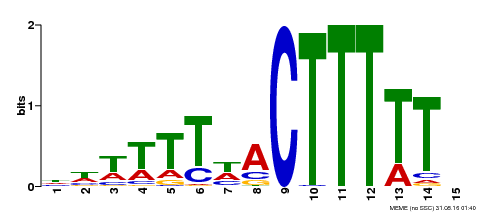
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cytokinin in procambium. Antagonized by the HD-ZIP III proteins and by mobile miR165 and miR166 microRNAs. {ECO:0000269|PubMed:30626969}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681841 | 0.0 | LN681841.1 Cucumis melo genomic scaffold, anchoredscaffold00011. | |||
| GenBank | LN713258 | 0.0 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008442559.1 | 0.0 | PREDICTED: dof zinc finger protein DOF5.1-like isoform X1 | ||||
| Swissprot | O80928 | 2e-52 | DOF24_ARATH; Dof zinc finger protein DOF2.4 | ||||
| TrEMBL | A0A1S3B5Z3 | 0.0 | A0A1S3B5Z3_CUCME; dof zinc finger protein DOF5.1-like isoform X1 | ||||
| STRING | XP_008442559.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF656 | 34 | 143 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G37590.1 | 3e-50 | DNA binding with one finger 2.4 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



